Repeated measures ANOVA
e.g. 1
demo1 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo1.csv")
## Convert variables to factor
demo1 <- within(demo1, {
group <- factor(group)
time <- factor(time)
id <- factor(id)
})
demo1
par(cex = .6)
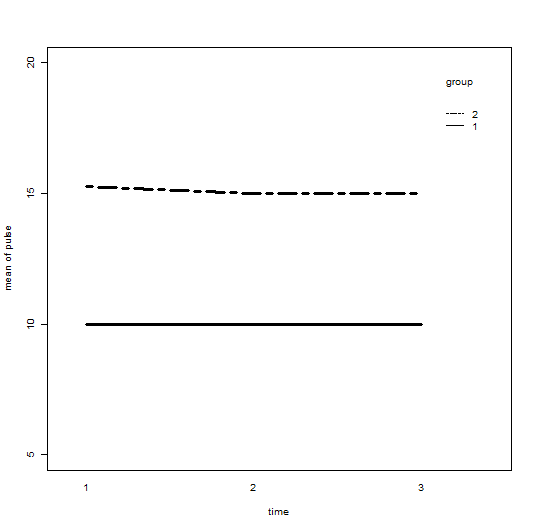
with(demo1, interaction.plot(time, group, pulse,
ylim = c(5, 20), lty= c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "time", trace.label = "group"))
demo1.aov <- aov(pulse ~ group * time + Error(id), data = demo1)
summary(demo1.aov)
>
> demo1 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo1.csv")
> ## Convert variables to factor
> demo1 <- within(demo1, {
+ group <- factor(group)
+ time <- factor(time)
+ id <- factor(id)
+ })
>
> demo1
id group pulse time
1 1 1 10 1
2 1 1 10 2
3 1 1 10 3
4 2 1 10 1
5 2 1 10 2
6 2 1 10 3
7 3 1 10 1
8 3 1 10 2
9 3 1 10 3
10 4 1 10 1
11 4 1 10 2
12 4 1 10 3
13 5 2 15 1
14 5 2 15 2
15 5 2 15 3
16 6 2 15 1
17 6 2 15 2
18 6 2 15 3
19 7 2 16 1
20 7 2 15 2
21 7 2 15 3
22 8 2 15 1
23 8 2 15 2
24 8 2 15 3
> par(cex = .6)
>
> with(demo1, interaction.plot(time, group, pulse,
+ ylim = c(5, 20), lty= c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "time", trace.label = "group"))
>
> demo1.aov <- aov(pulse ~ group * time + Error(id), data = demo1)
> summary(demo1.aov)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
group 1 155.04 155.04 3721 1.3e-09 ***
Residuals 6 0.25 0.04
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Error: Within
Df Sum Sq Mean Sq F value Pr(>F)
time 2 0.0833 0.04167 1 0.397
group:time 2 0.0833 0.04167 1 0.397
Residuals 12 0.5000 0.04167
>
>
e.g. 2
demo2 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo2.csv")
## Convert variables to factor
demo2 <- within(demo2, {
group <- factor(group)
time <- factor(time)
id <- factor(id)
})
demo2
par(cex = .6)
with(demo2, interaction.plot(time, group, pulse,
ylim = c(10, 40), lty = c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "time", trace.label = "group"))
demo2.aov <- aov(pulse ~ group * time + Error(id), data = demo2)
summary(demo2.aov)

>
> demo2 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo2.csv")
> ## Convert variables to factor
> demo2 <- within(demo2, {
+ group <- factor(group)
+ time <- factor(time)
+ id <- factor(id)
+ })
>
> demo2
id group pulse time
1 1 1 14 1
2 1 1 19 2
3 1 1 29 3
4 2 1 15 1
5 2 1 25 2
6 2 1 26 3
7 3 1 16 1
8 3 1 16 2
9 3 1 31 3
10 4 1 12 1
11 4 1 24 2
12 4 1 32 3
13 5 2 10 1
14 5 2 21 2
15 5 2 24 3
16 6 2 17 1
17 6 2 26 2
18 6 2 35 3
19 7 2 19 1
20 7 2 22 2
21 7 2 32 3
22 8 2 15 1
23 8 2 23 2
24 8 2 34 3
>
> par(cex = .6)
>
> with(demo2, interaction.plot(time, group, pulse,
+ ylim = c(10, 40), lty = c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "time", trace.label = "group"))
>
> demo2.aov <- aov(pulse ~ group * time + Error(id), data = demo2)
> summary(demo2.aov)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
group 1 15.04 15.04 0.836 0.396
Residuals 6 107.92 17.99
Error: Within
Df Sum Sq Mean Sq F value Pr(>F)
time 2 978.2 489.1 53.684 1.03e-06 ***
group:time 2 1.1 0.5 0.059 0.943
Residuals 12 109.3 9.1
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
e.g. 3
demo3 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo3.csv")
## Convert variables to factor
demo3 <- within(demo3, {
group <- factor(group)
time <- factor(time)
id <- factor(id)
})
demo3
par(cex = .6)
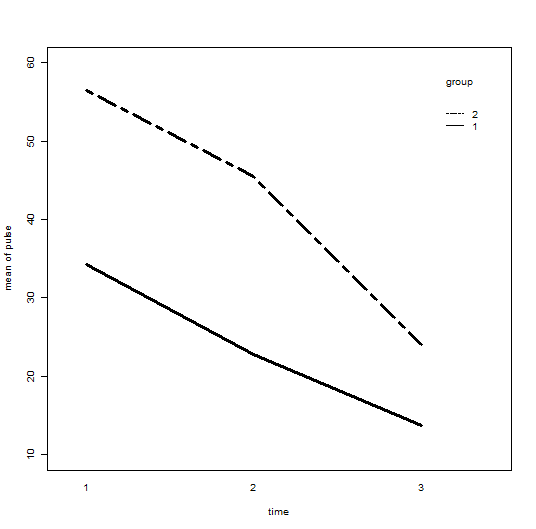
with(demo3, interaction.plot(time, group, pulse,
ylim = c(10, 60), lty = c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "time", trace.label = "group"))
demo3.aov <- aov(pulse ~ group * time + Error(id), data = demo3)
summary(demo3.aov)
> demo3 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo3.csv")
> ## Convert variables to factor
> demo3 <- within(demo3, {
+ group <- factor(group)
+ time <- factor(time)
+ id <- factor(id)
+ })
>
> demo3
id group pulse time
1 1 1 35 1
2 1 1 25 2
3 1 1 16 3
4 2 1 32 1
5 2 1 23 2
6 2 1 12 3
7 3 1 36 1
8 3 1 22 2
9 3 1 14 3
10 4 1 34 1
11 4 1 21 2
12 4 1 13 3
13 5 2 57 1
14 5 2 43 2
15 5 2 22 3
16 6 2 54 1
17 6 2 46 2
18 6 2 26 3
19 7 2 55 1
20 7 2 46 2
21 7 2 23 3
22 8 2 60 1
23 8 2 47 2
24 8 2 25 3
>
> par(cex = .6)
>
> with(demo3, interaction.plot(time, group, pulse,
+ ylim = c(10, 60), lty = c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "time", trace.label = "group"))
>
> demo3.aov <- aov(pulse ~ group * time + Error(id), data = demo3)
> summary(demo3.aov)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
group 1 2035.0 2035.0 343.1 1.6e-06 ***
Residuals 6 35.6 5.9
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Error: Within
Df Sum Sq Mean Sq F value Pr(>F)
time 2 2830.3 1415.2 553.8 1.52e-12 ***
group:time 2 200.3 100.2 39.2 5.47e-06 ***
Residuals 12 30.7 2.6
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
e.g. 4
demo4 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo4.csv")
## Convert variables to factor
demo4 <- within(demo4, {
group <- factor(group)
time <- factor(time)
id <- factor(id)
})
demo4
par(cex = .6)
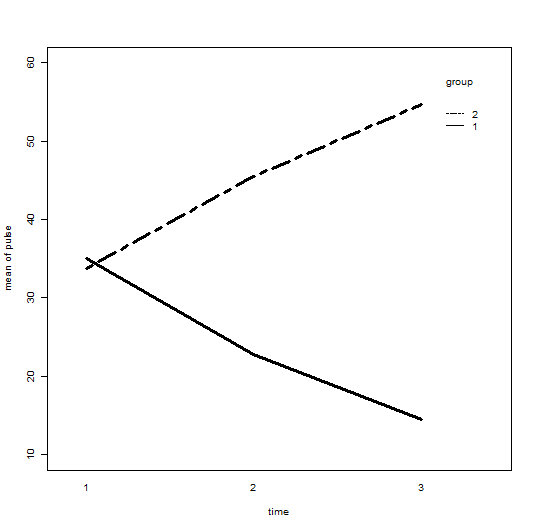
with(demo4, interaction.plot(time, group, pulse,
ylim = c(10, 60), lty = c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "time", trace.label = "group"))
demo4.aov <- aov(pulse ~ group * time + Error(id), data = demo4)
summary(demo4.aov)
>
> demo4 <- read.csv("https://stats.idre.ucla.edu/stat/data/demo4.csv")
> ## Convert variables to factor
> demo4 <- within(demo4, {
+ group <- factor(group)
+ time <- factor(time)
+ id <- factor(id)
+ })
>
> demo4
id group pulse time
1 1 1 35 1
2 1 1 25 2
3 1 1 12 3
4 2 1 34 1
5 2 1 22 2
6 2 1 13 3
7 3 1 36 1
8 3 1 21 2
9 3 1 18 3
10 4 1 35 1
11 4 1 23 2
12 4 1 15 3
13 5 2 31 1
14 5 2 43 2
15 5 2 57 3
16 6 2 35 1
17 6 2 46 2
18 6 2 58 3
19 7 2 37 1
20 7 2 48 2
21 7 2 51 3
22 8 2 32 1
23 8 2 45 2
24 8 2 53 3
>
> par(cex = .6)
>
> with(demo4, interaction.plot(time, group, pulse,
+ ylim = c(10, 60), lty = c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "time", trace.label = "group"))
>
> demo4.aov <- aov(pulse ~ group * time + Error(id), data = demo4)
> summary(demo4.aov)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
group 1 2542.0 2542 629 2.65e-07 ***
Residuals 6 24.3 4
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Error: Within
Df Sum Sq Mean Sq F value Pr(>F)
time 2 1 0.5 0.079 0.925
group:time 2 1736 868.2 137.079 5.44e-09 ***
Residuals 12 76 6.3
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
exer1
exer1 <- read.csv("https://stats.idre.ucla.edu/stat/data/exer.csv")
str(exer1)
exer1 <- within(exer1, {
diet <- factor(diet)
exertype <- factor(exertype)
time <- factor(time)
id <- factor(id)
}) ## 이제 pulse만 제외하고 모두 factor로 변환된 데이터
str(exer1)
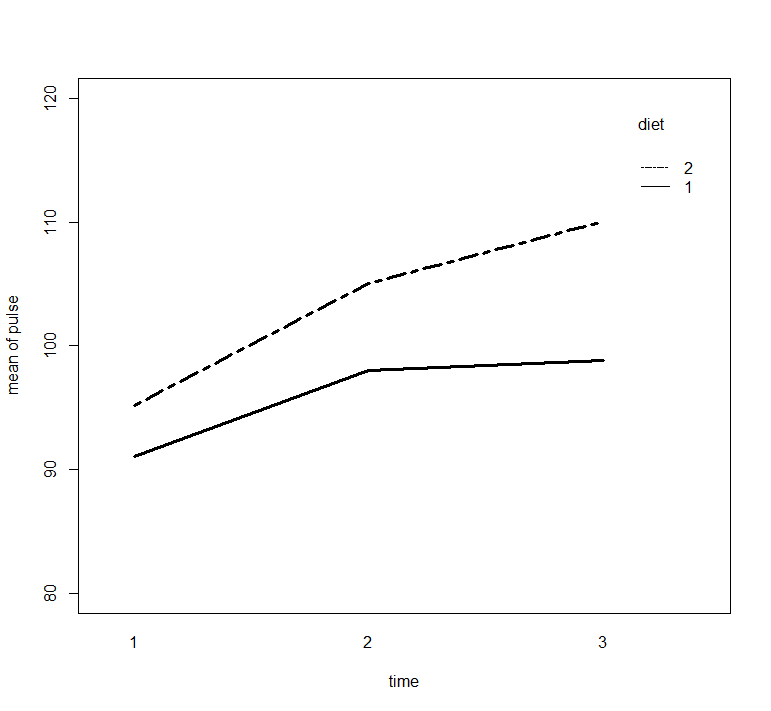
with(exer1, interaction.plot(time, diet, pulse,
ylim = c(80, 120), lty = c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "time", trace.label = "diet"))
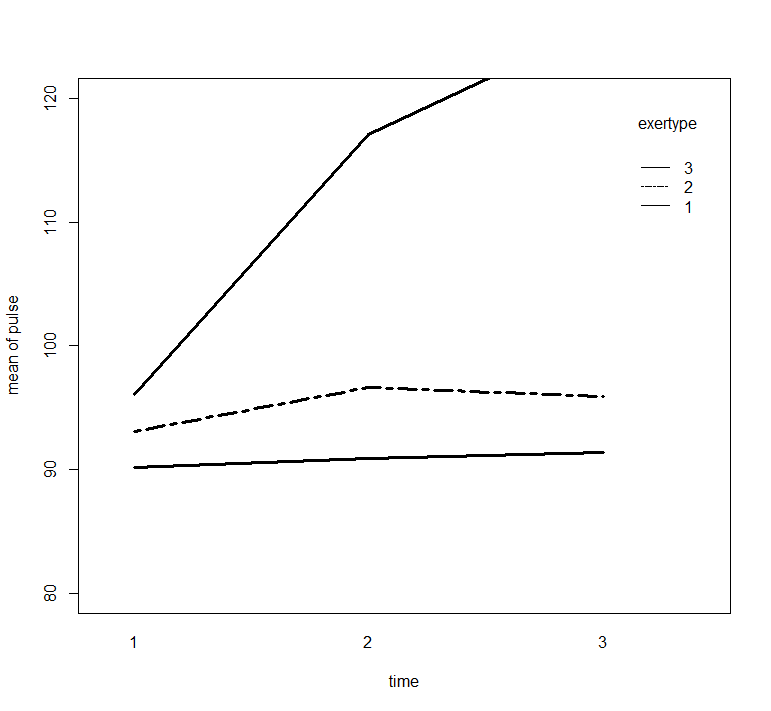
with(exer1, interaction.plot(time, exertype, pulse,
ylim = c(80, 120), lty = c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "time", trace.label = "exertype"))
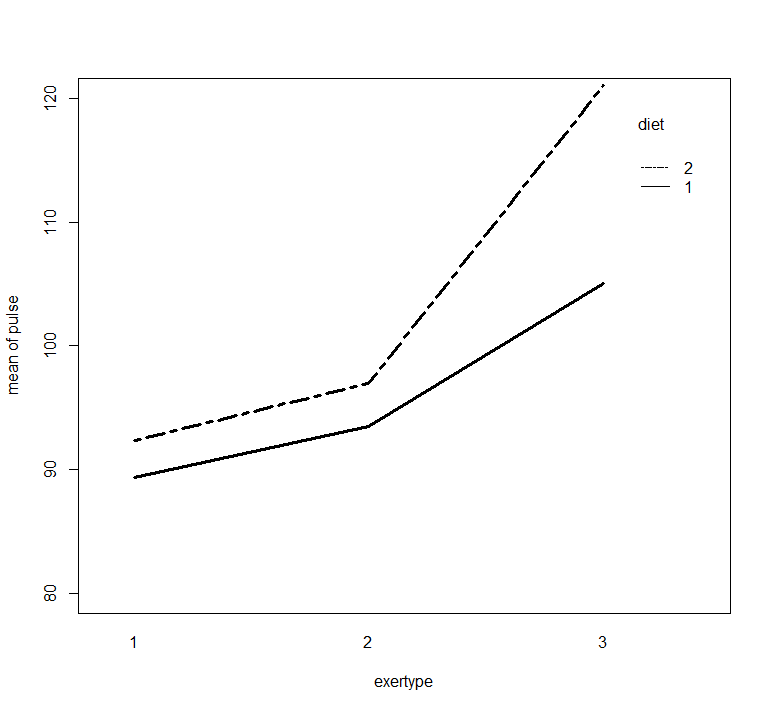
with(exer1, interaction.plot(exertype, diet, pulse,
ylim = c(80, 120), lty = c(1, 12), lwd = 3,
ylab = "mean of pulse", xlab = "exertype", trace.label = "diet"))
exer1a.aov <- aov(pulse ~ exertype * time + Error(id/time), data = exer1)
summary(exer1a.aov)
exer1b.aov <- aov(pulse ~ diet * time + Error(id/time), data = exer1)
summary(exer1b.aov)
exer1c.aov <- aov(pulse ~ diet * exertype, data = exer1)
summary(exer1c.aov)
> exer1 <- read.csv("https://stats.idre.ucla.edu/stat/data/exer.csv")
> str(exer1)
'data.frame': 90 obs. of 5 variables:
$ id : int 1 1 1 2 2 2 3 3 3 4 ...
$ diet : int 1 1 1 1 1 1 1 1 1 1 ...
$ exertype: int 1 1 1 1 1 1 1 1 1 1 ...
$ pulse : int 85 85 88 90 92 93 97 97 94 80 ...
$ time : int 1 2 3 1 2 3 1 2 3 1 ...
> exer1 <- within(exer1, {
+ diet <- factor(diet)
+ exertype <- factor(exertype)
+ time <- factor(time)
+ id <- factor(id)
+ }) ## 이제 pulse만 제외하고 모두 factor로 변환된 데이터
>
> str(exer1)
'data.frame': 90 obs. of 5 variables:
$ id : Factor w/ 30 levels "1","2","3","4",..: 1 1 1 2 2 2 3 3 3 4 ...
$ diet : Factor w/ 2 levels "1","2": 1 1 1 1 1 1 1 1 1 1 ...
$ exertype: Factor w/ 3 levels "1","2","3": 1 1 1 1 1 1 1 1 1 1 ...
$ pulse : int 85 85 88 90 92 93 97 97 94 80 ...
$ time : Factor w/ 3 levels "1","2","3": 1 2 3 1 2 3 1 2 3 1 ...
> with(exer1, interaction.plot(time, diet, pulse,
+ ylim = c(80, 120), lty = c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "time", trace.label = "diet"))
> with(exer1, interaction.plot(time, exertype, pulse,
+ ylim = c(80, 120), lty = c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "time", trace.label = "exertype"))
>
> with(exer1, interaction.plot(exertype, diet, pulse,
+ ylim = c(80, 120), lty = c(1, 12), lwd = 3,
+ ylab = "mean of pulse", xlab = "exertype", trace.label = "diet"))
>
>
> exer1a.aov <- aov(pulse ~ exertype * time + Error(id/time), data = exer1)
> summary(exer1a.aov)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
exertype 2 8326 4163 27 3.62e-07 ***
Residuals 27 4163 154
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Error: id:time
Df Sum Sq Mean Sq F value Pr(>F)
time 2 2067 1033.3 23.54 4.45e-08 ***
exertype:time 4 2723 680.8 15.51 1.65e-08 ***
Residuals 54 2370 43.9
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
> exer1b.aov <- aov(pulse ~ diet * time + Error(id/time), data = exer1)
> summary(exer1b.aov)
Error: id
Df Sum Sq Mean Sq F value Pr(>F)
diet 1 1262 1262 3.147 0.0869 .
Residuals 28 11227 401
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Error: id:time
Df Sum Sq Mean Sq F value Pr(>F)
time 2 2067 1033.3 11.808 5.26e-05 ***
diet:time 2 193 96.4 1.102 0.339
Residuals 56 4901 87.5
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
> exer1c.aov <- aov(pulse ~ diet * exertype, data = exer1)
> summary(exer1c.aov)
Df Sum Sq Mean Sq F value Pr(>F)
diet 1 1262 1262 11.465 0.00108 **
exertype 2 8326 4163 37.824 1.94e-12 ***
diet:exertype 2 816 408 3.706 0.02868 *
Residuals 84 9245 110
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
> exer1
id diet exertype pulse time
1 1 1 1 85 1
2 1 1 1 85 2
3 1 1 1 88 3
4 2 1 1 90 1
5 2 1 1 92 2
6 2 1 1 93 3
7 3 1 1 97 1
8 3 1 1 97 2
9 3 1 1 94 3
10 4 1 1 80 1
11 4 1 1 82 2
12 4 1 1 83 3
13 5 1 1 91 1
14 5 1 1 92 2
15 5 1 1 91 3
16 6 2 1 83 1
17 6 2 1 83 2
18 6 2 1 84 3
19 7 2 1 87 1
20 7 2 1 88 2
21 7 2 1 90 3
22 8 2 1 92 1
23 8 2 1 94 2
24 8 2 1 95 3
25 9 2 1 97 1
26 9 2 1 99 2
27 9 2 1 96 3
28 10 2 1 100 1
29 10 2 1 97 2
30 10 2 1 100 3
31 11 1 2 86 1
32 11 1 2 86 2
33 11 1 2 84 3
34 12 1 2 93 1
35 12 1 2 103 2
36 12 1 2 104 3
37 13 1 2 90 1
38 13 1 2 92 2
39 13 1 2 93 3
40 14 1 2 95 1
41 14 1 2 96 2
42 14 1 2 100 3
43 15 1 2 89 1
44 15 1 2 96 2
45 15 1 2 95 3
46 16 2 2 84 1
47 16 2 2 86 2
48 16 2 2 89 3
49 17 2 2 103 1
50 17 2 2 109 2
51 17 2 2 90 3
52 18 2 2 92 1
53 18 2 2 96 2
54 18 2 2 101 3
55 19 2 2 97 1
56 19 2 2 98 2
57 19 2 2 100 3
58 20 2 2 102 1
59 20 2 2 104 2
60 20 2 2 103 3
61 21 1 3 93 1
62 21 1 3 98 2
63 21 1 3 110 3
64 22 1 3 98 1
65 22 1 3 104 2
66 22 1 3 112 3
67 23 1 3 98 1
68 23 1 3 105 2
69 23 1 3 99 3
70 24 1 3 87 1
71 24 1 3 132 2
72 24 1 3 120 3
73 25 1 3 94 1
74 25 1 3 110 2
75 25 1 3 116 3
76 26 2 3 95 1
77 26 2 3 126 2
78 26 2 3 143 3
79 27 2 3 100 1
80 27 2 3 126 2
81 27 2 3 140 3
82 28 2 3 103 1
83 28 2 3 124 2
84 28 2 3 140 3
85 29 2 3 94 1
86 29 2 3 135 2
87 29 2 3 130 3
88 30 2 3 99 1
89 30 2 3 111 2
90 30 2 3 150 3
>






