Table of Contents
R script output
설명없는 full R script와 output은 r script and output
이미 우리는 샘플평균들 집합의 평균과 분산값이 무엇이 되는가를 수학적으로 알아보았다. mean and variance of the sample mean.
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
>
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
….
필요한 펑션
- rnorm2는 평균과 표준편차값을 (mean, sd) 갖는 n 개의 샘플을 랜덤하게 추출하는 것
- ss는 sum of square 값을 구하는 펑션
- ss는 ss/n-1 의 variance 혹은 mean square값을 구할 때의 분자부분으로
- error 혹은 residual의 제곱의 합이라고 부를 수 있다 (variance 참조)
> ################################
> N.p <- 1000000
> m.p <- 100
> sd.p <- 10
>
> p1 <- rnorm2(N.p, m.p, sd.p)
> mean(p1)
[1] 100
> sd(p1)
[1] 10
>
> p2 <- rnorm2(N.p, m.p+10, sd.p)
> mean(p2)
[1] 110
> sd(p2)
[1] 10
>
> m.p1 <- mean(p1)
> sd.p1 <- sd(p1)
> var(p1)
[,1]
[1,] 100
- rnorm2 펑션을 이용하여 p1과 p2 모집단을 구한다.
- 각 모집단의 평균은 100과 120이고, 표준편차는 모두 10이다.
- 모집단의 크기는 각각 백만이다.
pnorm
> hist(p1, breaks=50, col=rgb(0,.5,.5,.5)) > abline(v=mean(p1),lwd=2) > abline(v=mean(p1)-sd(p1), lwd=2) > abline(v=mean(p1)+sd(p1), lwd=2) > abline(v=c(m.p1-2*sd.p1, m.p1+2*sd.p1), lwd=2, col='red') > abline(v=c(m.p1-3*sd.p1, m.p1+3*sd.p1), lwd=2, col='green') > > # area bet black = 68% > # between red = 95% > # between green = 99% > > pnorm(m.p1+sd.p1, m.p1, sd.p1) [1] 0.8413447 > pnorm(m.p1-sd.p1, m.p1, sd.p1) [1] 0.1586553 > pnorm(m.p1+sd.p1, m.p1, sd.p1) - + pnorm(m.p1-sd.p1, m.p1, sd.p1) [1] 0.6826895 > > pnorm(m.p1+2*sd.p1, m.p1, sd.p1) - + pnorm(m.p1-2*sd.p1, m.p1, sd.p1) [1] 0.9544997 > > pnorm(m.p1+3*sd.p1, m.p1, sd.p1) - + pnorm(m.p1-3*sd.p1, m.p1, sd.p1) [1] 0.9973002 > > m.p1 [1] 100 > sd.p1 [1] 10 > > (m.p1-m.p1)/sd.p1 [1] 0 > ((m.p1-sd.p1) - m.p1) / sd.p1 [1] -1 > (120-100)/10 [1] 2 > pnorm(1)-pnorm(-1) [1] 0.6826895 > pnorm(2)-pnorm(-2) [1] 0.9544997 > pnorm(3)-pnorm(3) [1] 0 > > 1-pnorm(-2)*2 [1] 0.9544997 > pnorm(2)-pnorm(-2) [1] 0.9544997 > > pnorm(120, 100, 10) [1] 0.9772499 > pnorm(2)-pnorm(-2) [1] 0.9544997 > > > > > > >
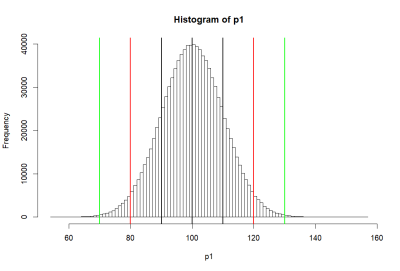
모집단 P1의 히스토그램 (histgram of P1)
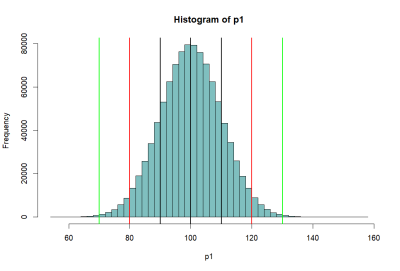
….
pnorm 펑션
- 위의 histogram에서 검정색의 선은 p1의 standard deviation값인 10씩 좌우로 그린것 (90과 110)
- 붉은 색선은 80, 120
- 녹색선은 70, 130 선을 말한다
- 고등학교 때, Normal distribution (정상분포의) 경우
- 평균을 중심으로 위아래로 SD값 하나씩 간 간격의 probability는 (면적은) 68%
- 두 개씩 간 값은 95%
- 세 개씩 간 값은 99% 라고 배웠다.
- pnorm은 percentage를 구하는 R의 명령어
- pnorm(m.p1+sd.p1, m.p1, sd.p1) 은
- 정규분포곡선에서
- 평균값과 표준편차값을 더한 값 (100+10=110) 을 기준으로
- 왼쪽의 부분이 몇 퍼센트인가를 구해주는 명령어
- m.p1+sd.p1은 110이므로
- 오른 쪽 검정색을 기준으로 왼쪽의 퍼센티지를 묻는 것
- 답은 0.8413447
- pnorm(m.p1-sd.p1, m.p1, sd.p1) - pnorm(90, 100, 10)이므로
- 왼쪽 검정색 선을 기준으로 왼쪽의 퍼센티지를 묻는 것
- 답은 0.1586553
- 전자에서 후자를 빼 준 값은 두개의 검정색 선의 안쪽 면적으로 묻는 것
- 답은 0.6826895
- 이 0.6826895 이 우리가 배운 68%
- 그렇다면 두 개씩 간 면적은 (붉은 색 선의 안쪽 부분)
- pnorm(m.p1+2*sd.p1, m.p1, sd.p1) - pnorm(m.p1-2*sd.p1, m.p1, sd.p1) = 0.9544997
- 0.9544997 혹은 95.44997%
- 이것이 우리가 배운 95%
- 마찬가지로 녹색선 가운데 부분은 pnorm(m.p1+3*sd.p1, m.p1, sd.p1) - pnorm(m.p1-3*sd.p1, m.p1, sd.p1) 로 구할 수 있는데 답은
- 0.9973002
- pnorm명령어는 pnorm(score, mean, sd)와 같이 사용하여 퍼센티지값을 구할 수 있는데
- pnorm(score)만으로 구하면, mean과 sd가 각각 0, 1을 default값으로 하고 생략이 된 것
- 위의 p1 모집단도 두번째 방법으로 구해 볼 수 있는데, 그렇게 하기 위해서
- p1을 표준점수화 하면 됨
- 표준점수화 한다는 뜻은 p1집합의 평균과 표준편차 값을 0과 1로 만든다는 것
- p1 모든 원소를 표준점수화하는 것은 각 점수와 평균점수 간의 차이에 sd가 몇개나 들어가는 지 구하는 것. 즉,
- $ \dfrac {(\text{score} - \text{m.p1})} {\text{sd.p1}} $
- 100점은 표준점수로 0이 된다 $ (100-100) / 10 = 0 $
- 110점은 1
- 90점은 -1
- 115점은 1.5
- 130점은 30/10 = 3 점으로 계산될 수 있다.
- 그렇다면 위의 pnorm(m.p1+sd.p1, m.p1, sd.p1)은 pnorm(1)과 같은 것
- pnorm(110, 100, 10) 이고 이 때 110점은 표준점수로 1 이다.
- 따라서 위의 68%, 95%, 99%는 pnorm(1)-pnorm(-1)과 같이 구할 수 있다.
z score, 표준점수
> za.p1 <- (p1-mean(p1))/sd(p1)
> zb.p1 <- scale(p1)
> tail(za.p1 == zb.p1, 10)
[,1]
[999991,] TRUE
[999992,] TRUE
[999993,] TRUE
[999994,] TRUE
[999995,] TRUE
[999996,] TRUE
[999997,] TRUE
[999998,] TRUE
[999999,] TRUE
[1000000,] TRUE
>
> mean(za.p1)
[1] -2.100958e-17
> mean(za.p1)==mean(zb.p1)
[1] TRUE
> round(mean(za.p1),10)
[1] 0
> sd(za.p1)
[1] 1
>
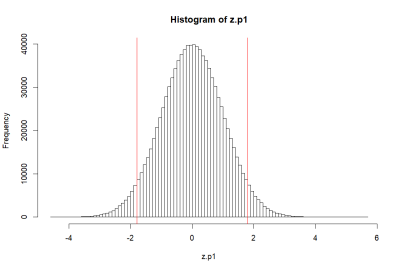
> pnorm(1.8)-pnorm(-1.8)
[1] 0.9281394
>
> hist(za.p1, breaks=50, col=rgb(1,0,0,0))
> abline(v=c(m.p1, -1.8, 1.8), col='red')
> 1-(pnorm(1.8)-pnorm(-1.8))
[1] 0.07186064
>
> > pnorm(1)-pnorm(-1) [1] 0.6826895 > 1-(pnorm(-1)*2) [1] 0.6826895 > > pnorm(2)-pnorm(-2) [1] 0.9544997 > 1-(pnorm(-2)*2) [1] 0.9544997 > > 1-(pnorm(-3)*2) [1] 0.9973002 >
….
qnorm
> # > hist(p1, breaks=50, col=rgb(.9,.9,.9,.9)) > abline(v=mean(p1),lwd=2) > abline(v=mean(p1)-sd(p1), lwd=2) > abline(v=mean(p1)+sd(p1), lwd=2) > abline(v=c(m.p1-2*sd.p1, m.p1+2*sd.p1), lwd=2, col='red') > abline(v=c(m.p1-3*sd.p1, m.p1+3*sd.p1), lwd=2, col='green') > > # 68% > a <- qnorm(.32/2) > b <- qnorm(1-.32/2) > c(a, b) [1] -0.9944579 0.9944579 > c(-1, 1) [1] -1 1 > # note that > .32/2 [1] 0.16 > pnorm(-1) [1] 0.1586553 > qnorm(.32/2) [1] -0.9944579 > qnorm(pnorm(-1)) [1] -1 > > # 95% > c <- qnorm(.05/2) > d <- qnorm(1-.05/2) > c(c, d) [1] -1.959964 1.959964 > c(-2,2) [1] -2 2 > > # 99% > e <- qnorm(.01/2) > f <- qnorm(1-.01/2) > c(e,f) [1] -2.575829 2.575829 > c(-3,3) [1] -3 3 > > > pnorm(b)-pnorm(a) [1] 0.68 > c(a, b) [1] -0.9944579 0.9944579 > pnorm(d)-pnorm(c) [1] 0.95 > c(c, d) [1] -1.959964 1.959964 > pnorm(f)-pnorm(e) [1] 0.99 > c(e, f) [1] -2.575829 2.575829 > > qnorm(.5) [1] 0 > qnorm(1) [1] Inf > >
qnorm
- qnorm는 pnorm의 반대값을 구하는 명령어
- 히스토그램에서 검정 색 부분의 바깥 쪽 부분은 32%이고 왼 쪽의 것은 이것의 반인 16% 이다.
- 이 16% 에 해당하는 표준점수가 무엇인가를 묻는 질문이
qnorm(.32/2)혹은qnorm(.32/2, 0, 1) - 원점수일 경우에 대입해서 물어본다면
qnorm(.32/2, 100, 10)과 같이 하면 된다 (혹은 qnorm(.32/2, m.p1, sd.p1) ). - 위의 명령어에 대한 답은 우리가 가장 쉽게 이해한 방법에 의하면
- -1 혹은 90 이렇게 될 것이다.
- 하지만 위에서 봤던 것처럼 pnorm(1)-pnorm(-1)이 정확히
- 0.6826895 인 것처럼, 우리가 이해한 방법은 정확한 값을 구해주지는 않는다.
- 마찬가지로 qnorm(0.05/2)와 qnorm(1-(0.05/2))는
- 표준편차 값인 1이 왼쪽으로 두개 내려가고 오른 쪽으로 두개 올라간 값의 quotient값을 구하는 것이므로
- 즉, 위의 histogram에서 붉은 색 부분의 바깥 쪽 면적에 해당하는 점수를 물어보는 것이므로
- -2와 2라고 대답해도 되지만 정확한 답은 아래와 같다 (1.96).
> qnorm(0.05/2) [1] -1.959964 > qnorm(1-(0.05/2)) [1] 1.959964 >
- 이에 해당하는 원점수 (p1 원소에 해당하는) 값은 80과 120이 아니라 아래와 같을 것이다.
> qnorm(0.05/2, 100, 10) [1] 80.40036 > qnorm(1-(0.05/2), 100, 10) [1] 119.5996 >
pnorm(110, 100, 10, lower.tail = F) pnorm((110-100)/10, lower.tail = F) pnorm(1, lower.tail = F) 1-pnorm(1) pnorm(-1) 1-(pnorm(-1)*2)
distribution of sample means
아래는 모두 같은 의미이다.
- distribution of sample means
- distribution of the sample mean
- mean and variance of the sample mean
- mean and variance of sample means
- sampling distribution
> ################################ > s.size <- 10 > > means <- c() > s1 <- sample(p1, s.size, replace = T) > mean(s1) [1] 100.7771 > means <- append(means, mean(s1)) > means <- append(means, mean(sample(p1, s.size, replace = T))) > means <- append(means, mean(sample(p1, s.size, replace = T))) > means <- append(means, mean(sample(p1, s.size, replace = T))) > means <- append(means, mean(sample(p1, s.size, replace = T))) > means [1] 100.77711 99.34268 101.12569 99.03624 98.73117 > hist(means) > > > > > > > > >
> ################################
> s.size <- 10
>
> means <- c()
> s1 <- sample(p1, s.size, replace = T)
> mean(s1)
[1] 100.7648
> means <- append(means, mean(s1))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
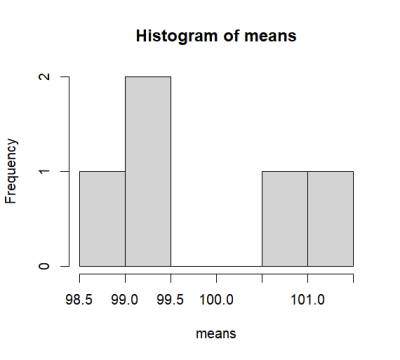
> means
[1] 100.7648 105.3139 107.5766 102.4201 101.6467
> hist(means)
>
> iter <- 100000
> # means <- c()
> means <- rep(NA, iter)
> tail(means)
[1] NA NA NA NA NA NA
>
> for (i in 1:iter) {
+ # means <- append(means, mean(sample(p1, s.size, replace = T)))
+ means[i] <- mean(sample(p1, s.size, replace = T))
+ }
> length(means)
[1] 100000
> mean(means)
[1] 100.0025
> sd(means)
[1] 3.167479
> var(means)
[1] 10.03292
> se.s <- sd(means)
>
….
- 이 집합의 평균과 분산을
mean(means),sd(means),var(means)으로 알아 보았는데 각각의 값이
> mean(means) [1] 100.0025 > sd(means) [1] 3.167479 > var(means) [1] 10.03292
- 위와 같다. 이 값은 샘플평균들의 평균이 무엇인가와 (mean(means))
- 샘플평균들의 분산값이 무엇인가를 (mean(means)) 구한것이고, 이는
- 우리가 이미 수학적으로 알아본 것과 같은 것이다 mean and variance of the sample mean
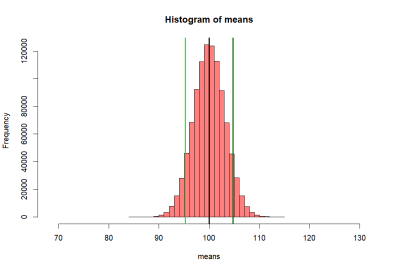
> # now we want to get sd of this distribution
> lo1 <- mean(means)-se.s
> hi1 <- mean(means)+se.s
> lo2 <- mean(means)-2*se.s
> hi2 <- mean(means)+2*se.s
> lo3 <- mean(means)-3*se.s
> hi3 <- mean(means)+3*se.s
> abline(v=mean(means), col="black", lwd=2)
> # abline(v=mean(p2), colo='darkgreen', lwd=2)
> abline(v=c(lo1, hi1, lo2, hi2, lo3, hi3),
+ col=c("red","red", "blue", "blue", "orange", "orange"),
+ lwd=2)
>
>
>
>
>
>
>
>
>
>
>
>
> # meanwhile . . . .
> se.s
[1] 3.167479
> sd(means)
[1] 3.167479
> var(means)
[1] 10.03292
>
> se.z <- sqrt(var(p1)/s.size)
> se.z
[,1]
[1,] 3.162278
> var.z <- var(p1)/s.size
> var.z
[,1]
[1,] 10
>
> # sd of sample means (sd(means))
> # = se.s
>
> # when iter value goes to
> # infinite value:
> # mean(means) = mean(p1)
> # and
> # var(means) = var(p1) / s.size
> # sd(means) = sqrt(var(p1)/s.size)
> # that is, se.s = se.z
> # This is called CLT (Central Limit Theorem)
> # see http://commres.net/central_limit_theorem
>
> mean(means)
[1] 100.0025
> var(means)
[1] 10.03292
> sd(means)
[1] 3.167479
> # remember we started talking sample size 10
> mean(p1)
[1] 100
> var(p1)/s.size
[,1]
[1,] 10
> sqrt(var(p1)/s.size)
[,1]
[1,] 3.162278
>
> # because CLT
> loz1 <- mean(p1)-se.z
> hiz1 <- mean(p1)+se.z
> loz2 <- mean(p1)-2*se.z
> hiz2 <- mean(p1)+2*se.z
> loz3 <- mean(p1)-3*se.z
> hiz3 <- mean(p1)+3*se.z
>
> c(lo1, loz1)
[1] 96.83502 96.83772
> c(lo2, loz2)
[1] 93.66754 93.67544
> c(lo3, loz3)
[1] 90.50006 90.51317
>
> c(hi1, hiz1)
[1] 103.1700 103.1623
> c(hi2, hiz2)
[1] 106.3375 106.3246
> c(hi3, hiz3)
[1] 109.5049 109.4868
>
>
>
>
mean and variance of the sample mean
….
- 샘플들을 직접 모아서 구한 샘플평균들의 표준편차 값은 (standard deviation of sample means, se.s = 3.161886)
- se.z 를 구해서 얻은 값과 같다 (3.162278).
> se.z <- sqrt(var(p1)/s.size) > se.z <- c(se.z) > se.z [1] 3.162278
- 사실, 우리가 백만 번의 샘플을 취해서 구한 means 집합의 평균과 표준편차 값은
- 이것을 정확하게는 mean of the distribution of sample means,
- standard deviation of the distribution of sample means 라고 부른다
- 그리고 standard deviation of the distribution of sample means를 흔히
- standard error라고 부른다.
- script에서 se.s 그리고 se.z 와 같은 se를 쓴 이유도 standard error 값을 구한 것이기 때문이다.
- 만약에 백만 번이 아니라 무한 대로 더 큰 숫자를 사용한다고 하면
- 위의 se.z 값을 구하는 식의 값을 갖게 된다. 이것을 말로 풀어서 설명하면
- 샘플평균들의 집합에서 표준편차 값은
- 원래 모집단의 분산값을 샘플사이즈로 나누어준 값에 제곱근을 씌워서 구할 수 있다이다 (아래 Latex 정리에서 3번)
\begin{eqnarray*} \overline{X} \sim \displaystyle \text{N} \left(\mu, \dfrac{\sigma^{2}}{n} \right) \end{eqnarray*}
\begin{eqnarray} & & \text{Normal distribution of sample means.} \\ & & \text{E[}\overline{X}\text{]} = \mu_{\overline{X}} = \mu \\ & & \text{V[}\overline{X}\text{]} = (\sigma_{\overline{X}})^2 = \frac{\sigma^2}{n} \;\; \text{or} \;\; \sigma_{\overline{X}} = \frac{\sigma}{\sqrt{n}} \end{eqnarray}
see Central Limit Theorem and Sampling Distribution
- 즉, 샘플평균을 모은 집합의 분산값은 그 샘플이 추출된 원래 population의 분산값을 샘플크기로 (sample size) 나누어 준 값이다.
- 즉,
var(means) = var(p1) / s.size - 따라서,
std(means) = sqrt(var(p1) / s.size) - 더하여 그 샘플평균 집합의 평균 값은 population의 평균값이 된다
- 즉,
mean(means) = mean(p1)
- 따라서 lo, hi에 해당하는 means분포의 값을 mean(means) +- sd(means)로 구했었는데,
- 샘플평균의 분포를 무한대 번을 했다고 하면 사실 이 값은
- mean(p1) +- se.z 로 구하는 것이 정확할 것이다.
- 여기서
se.z = sqrt(var(p1)/s.size)) - loz1 - hiz1, loz2 - hiz2 값들은 이렇게 구한 값들이다.
참고
> lo1 <- mean(means)-se.s > hi1 <- mean(means)+se.s > lo2 <- mean(means)-2*se.s > hi2 <- mean(means)+2*se.s > lo3 <- mean(means)-3*se.s > hi3 <- mean(means)+3*se.s
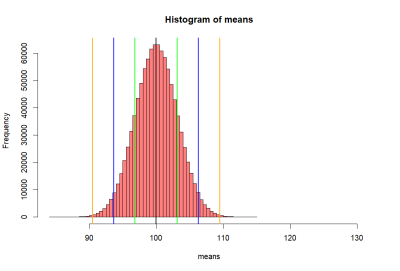
> means.10 <- rnorm2(iter, mean(p1), c(se.z))
> hist(means.10, breaks=50,
+ xlim = c(mean(p1)-5*se.z, mean(p1)+10*se.z),
+ col = rgb(1, 1, 1, .5))
> abline(v=mean(p1), col="black", lwd=3)
> # abline(v=mean(p2), colo='darkgreen', lwd=3)
> abline(v=c(loz1, hiz1, loz2, hiz2, loz3, hiz3),
+ col=c("darkgreen","darkgreen", "blue", "blue", "orange", "orange"),
+ lwd=2)
>
> round(c(lo1, hi1))
[1] 97 103
> round(c(lo2, hi2))
[1] 94 106
> round(c(lo3, hi3))
[1] 91 109
>
> round(c(loz1, hiz1))
[1] 97 103
> round(c(loz2, hiz2))
[1] 94 106
> round(c(loz3, hiz3))
[1] 91 109
>
>
>
Hypothesis test
> m.sample.i.got <- mean(means)+ 1.5*sd(means)
> m.sample.i.got
[1] 104.725
>
> hist(means.10, breaks=30,
+ xlim = c(mean(p1)-7*se.z, mean(p1)+10*se.z),
+ col = rgb(1, 1, 1, .5))
> abline(v=mean(p1), col="black", lwd=3)
> abline(v=m.sample.i.got, col='darkgreen', lwd=3)
>
> # what is the probablity of getting
> # greater than
> # m.sample.i.got?
> m.sample.i.got
[1] 104.725
> pnorm(m.sample.i.got, mean(p1), se.z, lower.tail = F)
[1] 0.06756624
>
> # then, what is the probabilty of getting
> # greater than m.sample.i.got and
> # less than corresponding value, which is
> # mean(means) - m.sample.i.got - mean(means)
> # (green line)
> tmp <- mean(p1) - (m.sample.i.got - mean(p1))
> tmp
[1] 95.27504
> abline(v=tmp, col='red', lwd=3)
> m.sample.i.got
[1] 104.725
> # 붉은 선의 왼쪽 부분을 구해서 2를 곱한 값의 범위
> prob.of.m.sample.i.got <- 2 * pnorm(m.sample.i.got, mean(p1), se.z, lower.tail = F)
> prob.of.m.sample.i.got
[1] 0.1351325
>
> (m.sample.i.got - mean(p1))/se.z
[,1]
[1,] 1.494165
> z.score <- (m.sample.i.got - mean(p1))/se.z
> pnorm(z.score, lower.tail = FALSE)
[,1]
[1,] 0.06756624
> 2 * pnorm(z.score, lower.tail = FALSE)
[,1]
[1,] 0.1351325
>
> # 즉 n = ssize 크기의 샘플을 구했을 때
> # 104.7489 점 오른쪽 그리고,
> # 이에 대응하는 95.27504 지점 왼쪽의
> # 평균이 나오는 probability는
> # 0.1351325
>
….
- 만약에 내가 한 샘플을 취해서 평균값을 살펴보니
- m.sample.i.got 값이었다고 하자 (104.725).
- 이 값보다 큰 값이거나 아니면
- 이 값에 해당하는 평균 반대편 값보다 작은 값이 값이
- 나올 확률은 무엇인가?
- 즉, 녹색선과 연두색 선 바깥 쪽 부분의 probability 값은?
- 아래처럼 구해서 13.5% 정도가 된다
- 즉, 모집단 p1에서
- 무작위로 샘플링을 하여 (see sampling and probability sampling)
- s.size의 (10) 샘플을 취했는데 그 샘플의 평균이
- 104.725 점보다 크거나 혹은
- 95.27504 점보다 작을 확률은 13.5% 이다.
> 2 * pnorm(m.sample.i.got, mean(p1), sd(means), lower.tail = F) [1] 0.1339882
- 그런데 위의 pnorm 은 표준점수화 해서 생각할 수 있다.
> (m.sample.i.got - mean(p1))/se.z
[,1]
[1,] 1.494165
> z.score <- (m.sample.i.got - mean(p1))/se.z
> pnorm(z.score, lower.tail = FALSE)
[,1]
[1,] 0.06756624
> 2 * pnorm(z.score, lower.tail = FALSE)
[,1]
[1,] 0.1351325
- 위처럼 z score를 구해서 pnorm으로 probability를 보는 것을 z-test 라고 한다.
> ### one more time
> # this time, with a story
> # 한편 우리는 특정한 평균과 표준편차를 갖는
> # p2를 알고 있다 (110, 10)
> mean(p2)
[1] 110
> sd(p2)
[1] 10
> set.seed(101)
> sample.i.got.p2 <- sample(p2, s.size)
> sample.i.got.p2
[1] 106.63828 107.09795 108.08661 119.09268 103.38813 118.30613 111.49542
[8] 94.25277 115.38045 95.49686
>
> m.sample.i.got.p2 <- mean(sample.i.got.p2)
> m.sample.i.got.p2
[1] 107.9235
>
> # 위의 샘플 평균이 위에서 얘기했던 샘플평균들의 집합에서
> # 나올 확률은 무엇일까?
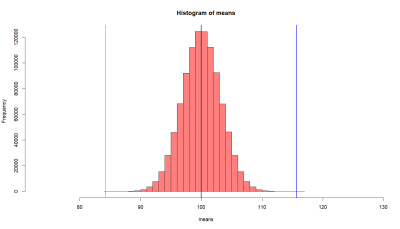
> hist(means.10, breaks=30,
+ xlim = c(tmp-10*se.z, m.sample.i.got+10*se.z),
+ col = rgb(1, 1, 1, .5))
> abline(v=mean(p1), col="black", lwd=3)
> abline(v=m.sample.i.got.p2, col='blue', lwd=3)
>
> # what is the probablity of getting
> # greater than
> # m.sample.i.got?
> m.sample.i.got.p2
[1] 107.9235
> pnorm(m.sample.i.got.p2, mean(p1), se.z, lower.tail = F)
[1] 0.006111512
> # 혹은
> z.cal <- (m.sample.i.got.p2-mean(p1))/se.z
> pnorm(z.cal, lower.tail = F)
[,1]
[1,] 0.006111512
>
> # then, what is the probability of getting
> # greater than m.sample.i.got and
> # less than corresponding value, which is
> # mean(means) - m.sample.i.got - mean(means)
> # (green line)
> abline(v=mean(p1)-(m.sample.i.got.p2-mean(p1)), col='red', lwd=3)
> 2 * pnorm(m.sample.i.got.p2, mean(p1), se.z, lower.tail = F)
[1] 0.01222302
> 2 * pnorm(z.cal, lower.tail = F)
[,1]
[1,] 0.01222302
>
> # 위는 샘플평균과 모집단 평균값 간의 차이 diff를
> # standard error값으로 즉,
> # sd of sample means (n=s.size) 값으로 나눠 준
> # 표준점수 값을 구하고
> # 이 점수에 대한 probability값을 구해 본것
>
> diff <- m.sample.i.got.p2 - mean(p1)
> se.z <- sqrt(var(p1)/s.size)
> z.cal <- diff / se.z
> z.cal
[,1]
[1,] 2.505639
> prob.z <- pnorm(z.cal, lower.tail=F)*2
> prob.z
[,1]
[1,] 0.01222302
> # 위에 대한 해석 . . . . .
> #
> #
> ###################################################
….
다른 모집단에서 온 샘플 (sample from other population, p2)
이전 처럼 10명을 샘플로 뽑는데, p2에서 뽑는다. 따라서 이 샘플의 평균은 107.9235 이다. 그런데 연구자는 이 샘플이 나온 모집단은 전혀 모르는 상태이다. 모집단의 평균도 모르고 분산값도 모른다. 그냥 10명을 샘플로 뽑았고, 이들이 p1에서 나왔을 확률을 알아보려고 한다. 이를 알아보기 위해서 스크립트를 돌려 보니, blue와 green라인 밖의 범위가 나올 확률은 0.01222302로 5/100 보다 작고, 1/100 에 가깝다.
이 probability level이 어느정도나 작아야 이 샘플이 p1에서 나오지 않고 p2에서 나왔다고 판단할 수 있을까? 관습적으로 5/100를 기준으로 해서 이 범위보다 작게 되면 p1의 모집단에서 나온 샘플이 아닌 것으로 판단하게 된다. 이 논리에 따라서, (평균을 107.9235 값을 갖는)이 샘플은, p1에서 나오기 힘든 샘플이라고 판단 된다.
- R에는 z-test 펑션이 없다. 현실에서는 전체 모집단의 평균을 알고 있는 경우는 많지만 표준편차까지 알고 있는 경우는 많지 않다. 그래서 많이 쓰이지 않는 편이다.
- 모집단의 평균과 표준편차를 알고 있다고 하면, 우리는 R에서 z test를 하는 절차는
- n = n 일 경우의 샘플링분포에서 se 를 구한다
- se = sigma / sqrt(n)
- 테스트할 점수의 z score를 구한다.
- diff = test.score - mean.of.population
- z.score = diff / se
- z score 보다 큰 점수나 -z score 보다 작은 점수가 나올 확률를 위의 샘플링 분포에 구한다.
- p.value = 2 * pnorm(z.score, lower.tail=F)
- z.score와 p.vallue로 테스트점수가 모집단에서 나왔는지 나올 수 없는지 (나오기 어려운지를) 판단한다.
> ###################################################
> # 다른 케이스 --> t-test 케이스
> # what if we do not know the sd of
> # population?
> # we use sd(sample) instead
> # obtained z-score is called t-score
> # and we use t-distribution to get
> # the probability of the t-score instead
> # of normal distribution (z).
> diff
[1] 7.923527
> se.t <- sqrt(var(sample.i.got.p2)/s.size)
> t.cal <- diff/se.t
> t.cal
[1] 2.914599
> pt(t.cal, df=s.size-1, lower.tail = F)
[1] 0.008591086
> pt(t.cal, df=s.size-1, lower.tail = F)*2
[1] 0.01718217
> prob.t <- pt(t.cal, df=s.size-1, lower.tail = F)*2
>
> # note that
> z.cal
[,1]
[1,] 2.505639
> se.z
[,1]
[1,] 3.162278
> prob.z
[,1]
[1,] 0.01222302
>
> t.cal
[1] 2.914599
> se.t
[1] 2.718566
> prob.t
[1] 0.01718217
>
> t.test(sample.i.got.p2, mu=mean(p1))
One Sample t-test
data: sample.i.got.p2
t = 2.9146, df = 9, p-value = 0.01718
alternative hypothesis: true mean is not equal to 100
95 percent confidence interval:
101.7737 114.0733
sample estimates:
mean of x
107.9235
t-test 중에서 2번째 케이스
- 모집단의 평균은 알지만 표준편차 정보는 없는 경우이다.
- 똑같은 논리로 생각을 해서 모집단의 샘플링분포를 (distribution of sample means) 머리에 두고
- se값을 구한다. 이 때의 se 값은
se.cal ← sqrt( s^2/n )값으로 구한다.- sigma 대신에 s를 사용한 것에 주목
- z.score에 해당하는 t.score를 구한다.
- 테스트점수와 모집단 평균의 차이를 구한 후 (
diff = test.score - mean(p1)) - se.cal 값으로 나눠준다.
t.cal ← diff / se.cal
- t.cal 값 이상, 반대편 점수의 이하가 나올 확률을 구한다.
- 이 때, 모집단의 표준편차를 사용해서 z.score를 구하지 않았으므로
- 그리고, 이 probability는 샘플의 크기 n에 영향을 받으므로 n의 크기에 따라서 변화하는 probability distribution을 사용한다.
- p.value ← pt(t.score, df=n-1, lower.tail=F) * 2
- t.cal과 p.value로 테스트점수가 나올 가능성을 판단하여 가설을 기각하거나 채택한다 (검증한다).
types of error
hypothesis testing
> # 그렇다면 > # 두 샘플의 평균과 표준편차만 알고 > # 모집단들의 파라미터는 모를 때는? > > # note that > # v(x)+v(y) = v(x)+v(y) > # = ss/dfx + ss/dfy > > s.size = 30 > s1 <- sample(p1, s.size, replace = T) > s2 <- sample(p2, s.size, replace = T) > s1 [1] 100.46855 97.67673 94.48868 97.51996 94.28300 113.78623 95.44617 [8] 92.91108 97.46601 100.66815 113.94452 81.20774 93.77529 93.64388 [15] 111.43702 104.34787 98.82403 100.93909 111.08048 110.12275 94.68502 [22] 100.14734 80.23712 88.02110 118.89602 99.46530 106.92325 97.13274 [29] 106.40479 107.44831 > s2 [1] 120.28360 105.80657 116.28974 106.63910 108.66632 107.51868 123.66436 [8] 96.06523 111.53770 108.78226 115.22628 104.60990 105.07035 114.90022 [15] 103.92860 105.06587 113.77416 108.07776 111.92824 112.01554 104.34549 [22] 100.51113 84.75464 96.36697 117.09381 116.29935 117.20402 109.61657 [29] 106.46650 105.86733 > > mean(s1) [1] 100.1133 > mean(s2) [1] 108.6125 > sd(s1) [1] 9.132472 > sd(s2) [1] 7.916677 > > ss(s1) [1] 2418.659 > ss(s2) [1] 1817.54 > df1 <- s.size - 1 > df2 <- s.size - 1 > v.pooled <- (ss(s1)+ss(s2))/(df1+df2) > se.tst <- sqrt((v.pooled/s.size)+(v.pooled)/s.size) > diff <- abs(mean(s2)-mean(s1)) > t.ts.cal <- diff/se.tst > t.ts.cal [1] 3.851705 > pt(t.ts.cal, df=df1+df2, lower.tail = F)*2 [1] 0.0002954557 > > t.test(s2, s1, var.equal=T) Two Sample t-test data: s2 and s1 t = 3.8517, df = 58, p-value = 0.0002955 alternative hypothesis: true difference in means is not equal to 0 95 percent confidence interval: 4.082229 12.916309 sample estimates: mean of x mean of y 108.6125 100.1133 >
t-test 중에서 3번째
R script and output
rs01
# http://commres.net/mean_and_variance_of_the_sample_mean
# we understand the concept of sampling distribution
rm(list=ls())
rnorm2 <- function(n,mean,sd){
mean+sd*scale(rnorm(n))
}
se <- function(sample) {
sd(sample)/sqrt(length(sample))
}
ss <- function(x) {
sum((x-mean(x))^2)
}
################################
N.p <- 1000000
m.p <- 100
sd.p <- 10
p1 <- rnorm2(N.p, m.p, sd.p)
mean(p1)
sd(p1)
p2 <- rnorm2(N.p, m.p+10, sd.p)
mean(p2)
sd(p2)
m.p1 <- mean(p1)
sd.p1 <- sd(p1)
var(p1)
hist(p1, breaks=50, col = rgb(1, 1, 1, 0.5),
main = "histogram of p1 and p2",)
abline(v=mean(p1), col="black", lwd=3)
hist(p2, add=T, breaks=50, col=rgb(1,1,.5,.5))
abline(v=mean(p2), col="red", lwd=3)
hist(p1, breaks=50, col=rgb(0,.5,.5,.5))
abline(v=mean(p1),lwd=2)
abline(v=mean(p1)-sd(p1), lwd=2)
abline(v=mean(p1)+sd(p1), lwd=2)
abline(v=c(m.p1-2*sd.p1, m.p1+2*sd.p1), lwd=2, col='red')
abline(v=c(m.p1-3*sd.p1, m.p1+3*sd.p1), lwd=2, col='green')
# area bet black = 68%
# between red = 95%
# between green = 99%
pnorm(m.p1+sd.p1, m.p1, sd.p1)
pnorm(m.p1-sd.p1, m.p1, sd.p1)
pnorm(m.p1+sd.p1, m.p1, sd.p1) -
pnorm(m.p1-sd.p1, m.p1, sd.p1)
pnorm(m.p1+2*sd.p1, m.p1, sd.p1) -
pnorm(m.p1-2*sd.p1, m.p1, sd.p1)
pnorm(m.p1+3*sd.p1, m.p1, sd.p1) -
pnorm(m.p1-3*sd.p1, m.p1, sd.p1)
m.p1
sd.p1
# 표준점수 (standardized score)
za.p1 <- (p1-mean(p1))/sd(p1)
# e.g.,
za.eg.1 <- (120 - 100) / 10
za.eg.1
# function
zb.p1 <- scale(p1)
tail(za.p1 == zb.p1, 10)
mean(za.p1)
mean(za.p1)==mean(zb.p1)
round(mean(za.p1),10)
sd(za.p1)
pnorm(110, 100, 10)
pnorm((110-100)/10)
pnorm(1)
pnorm(90, 100, 10)
pnorm((90-100)/10)
pnorm(-1)
pnorm(110, 100, 10) - pnorm(90, 100, 10)
pnorm(1) - pnorm(-1)
pnorm(120, 100, 10) - pnorm(80, 100, 10)
pnorm(2) - pnorm(-2)
pnorm(118, 100, 10)
pnorm(82, 100, 10)
pnorm(118, 100, 10) - pnorm(82, 100, 10)
pnorm(1.8)
pnorm(-1.8)
pnorm(1.8) - pnorm(-1.8)
hist(za.p1, breaks=50, col=rgb(1,0,0,0))
abline(v=c(m.p1, -1.8, 1.8), col='red')
1-(pnorm(1.8)-pnorm(-1.8))
pnorm(1)-pnorm(-1)
1-(pnorm(-1)*2)
pnorm(2)-pnorm(-2)
1-(pnorm(-2)*2)
1-(pnorm(-3)*2)
#
hist(p1, breaks=50, col=rgb(.9,.9,.9,.9))
abline(v=mean(p1),lwd=2)
abline(v=mean(p1)-sd(p1), lwd=2)
abline(v=mean(p1)+sd(p1), lwd=2)
abline(v=c(m.p1-2*sd.p1, m.p1+2*sd.p1), lwd=2, col='red')
abline(v=c(m.p1-3*sd.p1, m.p1+3*sd.p1), lwd=2, col='green')
# 68%
a <- qnorm(.32/2)
b <- qnorm(1-.32/2)
c(a, b)
c(-1, 1)
pnorm(a)
pnorm(b)
pnorm(-1)
pnorm(1)
# 95%
c <- qnorm(.05/2)
d <- qnorm(1-.05/2)
c(c, d)
c(-2,2)
# 99%
e <- qnorm(.01/2)
f <- qnorm(1-.01/2)
c(e,f)
c(-3,3)
pnorm(b)-pnorm(a)
c(a, b)
c(-1, 1)
pnorm(d)-pnorm(c)
c(c, d)
c(-2, 2)
pnorm(f)-pnorm(e)
c(e, f)
c(-3, 3)
qnorm(.5)
qnorm(1)
################################
s.size <- 10
means <- c()
s1 <- sample(p1, s.size, replace = T)
mean(s1)
means <- append(means, mean(s1))
means <- append(means, mean(sample(p1, s.size, replace = T)))
means <- append(means, mean(sample(p1, s.size, replace = T)))
means <- append(means, mean(sample(p1, s.size, replace = T)))
means <- append(means, mean(sample(p1, s.size, replace = T)))
means
hist(means)
iter <- 100000
# means <- c()
means <- rep(NA, iter)
tail(means)
for (i in 1:iter) {
# means <- append(means, mean(sample(p1, s.size, replace = T)))
means[i] <- mean(sample(p1, s.size, replace = T))
}
length(means)
mean(means)
sd(means)
var(means)
se.s <- sd(means)
hist(means, breaks=50,
xlim = c(mean(means)-5*sd(means), mean(means)+10*sd(means)),
col=rgb(1, 1, 1, .5))
abline(v=mean(means), col="black", lwd=3)
# now we want to get sd of this distribution
lo1 <- mean(means)-se.s
hi1 <- mean(means)+se.s
lo2 <- mean(means)-2*se.s
hi2 <- mean(means)+2*se.s
lo3 <- mean(means)-3*se.s
hi3 <- mean(means)+3*se.s
abline(v=mean(means), col="black", lwd=2)
# abline(v=mean(p2), colo='darkgreen', lwd=2)
abline(v=c(lo1, hi1, lo2, hi2, lo3, hi3),
col=c("red","red", "blue", "blue", "orange", "orange"),
lwd=2)
# meanwhile . . . .
se.s
sd(means)
var(means)
se.z <- sqrt(var(p1)/s.size)
se.z
var.z <- var(p1)/s.size
var.z
# sd of sample means (sd(means))
# = se.s
# when iter value goes to
# infinite value:
# mean(means) = mean(p1)
# and
# var(means) = var(p1) / s.size
# sd(means) = sqrt(var(p1)/s.size)
# that is, se.s = se.z
# This is called CLT (Central Limit Theorem)
# see http://commres.net/wiki/cetral_limit_theorem
mean(means)
var(means)
sd(means)
# remember we started talking sample size 10
mean(p1)
var(p1)/s.size
sqrt(var(p1)/s.size)
# because CLT
loz1 <- mean(p1)-se.z
hiz1 <- mean(p1)+se.z
loz2 <- mean(p1)-2*se.z
hiz2 <- mean(p1)+2*se.z
loz3 <- mean(p1)-3*se.z
hiz3 <- mean(p1)+3*se.z
c(lo1, loz1)
c(lo2, loz2)
c(lo3, loz3)
c(hi1, hiz1)
c(hi2, hiz2)
c(hi3, hiz3)
means.10 <- rnorm2(iter, mean(p1), c(se.z))
hist(means.10, breaks=50,
xlim = c(mean(p1)-5*se.z, mean(p1)+10*se.z),
col = rgb(1, 1, 1, .5))
abline(v=mean(p1), col="black", lwd=3)
# abline(v=mean(p2), colo='darkgreen', lwd=3)
abline(v=c(loz1, hiz1, loz2, hiz2, loz3, hiz3),
col=c("darkgreen","darkgreen", "blue", "blue", "orange", "orange"),
lwd=2)
round(c(lo1, hi1))
round(c(lo2, hi2))
round(c(lo3, hi3))
round(c(loz1, hiz1))
round(c(loz2, hiz2))
round(c(loz3, hiz3))
m.sample.i.got <- mean(means)+ 1.5*sd(means)
m.sample.i.got
hist(means.10, breaks=30,
xlim = c(mean(p1)-7*se.z, mean(p1)+10*se.z),
col = rgb(1, 1, 1, .5))
abline(v=mean(p1), col="black", lwd=3)
abline(v=m.sample.i.got, col='darkgreen', lwd=3)
# what is the probablity of getting
# greater than
# m.sample.i.got?
m.sample.i.got
pnorm(m.sample.i.got, mean(p1), se.z, lower.tail = F)
# then, what is the probabilty of getting
# greater than m.sample.i.got and
# less than corresponding value, which is
# mean(means) - m.sample.i.got - mean(means)
# (green line)
tmp <- mean(p1) - (m.sample.i.got - mean(p1))
tmp
abline(v=tmp, col='red', lwd=3)
m.sample.i.got
# 붉은 선의 왼쪽 부분을 구해서 2를 곱한 값의 범위
prob.of.m.sample.i.got <- 2 * pnorm(m.sample.i.got, mean(p1), se.z, lower.tail = F)
prob.of.m.sample.i.got
(m.sample.i.got - mean(p1))/se.z
z.score <- (m.sample.i.got - mean(p1))/se.z
pnorm(z.score, lower.tail = FALSE)
2 * pnorm(z.score, lower.tail = FALSE)
# 즉 n = ssize 크기의 샘플을 구했을 때
# 104.7489 점 오른쪽 그리고,
# 이에 대응하는 95.25113 지점 왼쪽의
# 평균이 나오는 probability는
# 0.1331684
### one more time
# this time, with a story
# 한편 우리는 특정한 평균과 표준편차를 갖는
# p2를 알고 있다 (110, 10)
mean(p2)
sd(p2)
set.seed(101)
sample.i.got.p2 <- sample(p2, s.size)
sample.i.got.p2
m.sample.i.got.p2 <- mean(sample.i.got.p2)
m.sample.i.got.p2
# 위의 샘플 평균이 위에서 얘기했던 샘플평균들의 집합에서
# 나올 확률은 무엇일까?
hist(means.10, breaks=30,
xlim = c(tmp-10*se.z, m.sample.i.got+10*se.z),
col = rgb(1, 1, 1, .5))
abline(v=mean(p1), col="black", lwd=3)
abline(v=m.sample.i.got.p2, col='blue', lwd=3)
# what is the probablity of getting
# greater than
# m.sample.i.got?
m.sample.i.got.p2
pnorm(m.sample.i.got.p2, mean(p1), se.z, lower.tail = F)
# 혹은
z.cal <- (m.sample.i.got.p2-mean(p1))/se.z
pnorm(z.cal, lower.tail = F)
# then, what is the probability of getting
# greater than m.sample.i.got and
# less than corresponding value, which is
# mean(means) - m.sample.i.got - mean(means)
# (green line)
abline(v=mean(p1)-(m.sample.i.got.p2-mean(p1)), col='red', lwd=3)
2 * pnorm(m.sample.i.got.p2, mean(p1), se.z, lower.tail = F)
2 * pnorm(z.cal, lower.tail = F)
# 위는 샘플평균과 모집단 평균값 간의 차이 diff를
# standard error값으로 즉,
# sd of sample means (n=s.size) 값으로 나눠 준
# 표준점수 값을 구하고
# 이 점수에 대한 probability값을 구해 본것
diff <- m.sample.i.got.p2 - mean(p1)
se.z <- sqrt(var(p1)/s.size)
z.cal <- diff / se.z
z.cal
prob.z <- pnorm(z.cal, lower.tail=F)*2
prob.z
# 위에 대한 해석 . . . . .
#
#
###################################################
###################################################
# 다른 케이스 --> t-test 케이스
# what if we do not know the sd of
# population?
# we use sd(sample) instead
# obtained z-score is called t-score
# and we use t-distribution to get
# the probability of the t-score instead
# of normal distribution (z).
diff
se.t <- sqrt(var(sample.i.got.p2)/s.size)
t.cal <- diff/se.t
t.cal
pt(t.cal, df=s.size-1, lower.tail = F)
pt(t.cal, df=s.size-1, lower.tail = F)*2
prob.t <- pt(t.cal, df=s.size-1, lower.tail = F)*2
# note that
z.cal
se.z
prob.z
t.cal
se.t
prob.t
t.test(sample.i.got.p2, mu=mean(p1))
# 그렇다면
# 두 샘플의 평균과 표준편차만 알고
# 모집단들의 파라미터는 모를 때는?
# note that
# v(x)+v(y) = v(x)+v(y)
# = ss/dfx + ss/dfy
s.size = 30
s1 <- sample(p1, s.size, replace = T)
s2 <- sample(p2, s.size, replace = T)
s1
s2
mean(s1)
mean(s2)
sd(s1)
sd(s2)
ss(s1)
ss(s2)
df1 <- s.size - 1
df2 <- s.size - 1
v.pooled <- (ss(s1)+ss(s2))/(df1+df2)
se.tst <- sqrt((v.pooled/s.size)+(v.pooled)/s.size)
diff <- abs(mean(s2)-mean(s1))
t.ts.cal <- diff/se.tst
t.ts.cal
pt(t.ts.cal, df=df1+df2, lower.tail = F)*2
t.test(s2, s1, var.equal=T)
comb <- list(s1 = s1, s2 = s2)
op <- stack(comb)
head(op)
colnames(op)[1] <- "values"
colnames(op)[2] <- "group"
op$group <- factor(op$group)
head(op)
boxplot(op$values~op$group)
# Calculate the mean of 'Value' grouped by 'Category'
m.by.group <- aggregate(values ~ group, data = op, FUN = mean, na.rm = TRUE)
m.by.group
m.s1 <- m.by.group$values[1]
m.s2 <- m.by.group$values[2]
m.tot <- mean(op$values)
m.s1 <- mean(s1)
m.s2 <- mean(s2)
n.s1 <- length(s1)
n.s2 <- length(s2)
df.s1 <- n.s1-1
df.s2 <- n.s2-1
ss.tot <- sum((m.tot-op$values)^2)
ss.tot
ss.tot <- ss(op$values)
ss.tot
ss.bet <- ((m.tot-m.s1)^2*n.s1) +
((m.tot-m.s2)^2*n.s2)
ss.wit <- ss(s1) + ss(s2)
ss.tot
ss.bet
ss.wit
ss.bet+ss.wit
df.tot <- (n.s1+n.s2)-1
df.bet <- nlevels(op$group)-1
df.wit <- df.s1+df.s2
ms.tot <- ss.tot / df.tot
ms.bet <- ss.bet / df.bet
ms.wit <- ss.wit / df.wit
r.sq <- ss.bet / ss.tot
r.sq
f.cal <- ms.bet / ms.wit
f.cal
prob.f <- pf(f.cal, df.bet, df.wit, lower.tail = F)
prob.f
ss.bet
df.bet
ms.bet
ss.wit
df.wit
ms.wit
f.cal
prob.f
m.f <- aov(values~group, data=op)
summary(m.f)
m.t <- t.test(s2, s1, var.equal = T)
m.t
m.t$statistic^2
m.t$p.value
m.l <- lm(values~group, data=op)
summary(m.l)
f.cal
df.bet
df.wit
prob.f
r.sq
anova(m.l)
ss.bet
ss.wit
ms.bet
ms.wit
f.cal
prob.f
ro01
> # http://commres.net/mean_and_variance_of_the_sample_mean
> # we understand the concept of sampling distribution
>
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
>
> se <- function(sample) {
+ sd(sample)/sqrt(length(sample))
+ }
>
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
> ################################
> N.p <- 1000000
> m.p <- 100
> sd.p <- 10
>
> p1 <- rnorm2(N.p, m.p, sd.p)
> mean(p1)
[1] 100
> sd(p1)
[1] 10
>
> p2 <- rnorm2(N.p, m.p+10, sd.p)
> mean(p2)
[1] 110
> sd(p2)
[1] 10
>
> m.p1 <- mean(p1)
> sd.p1 <- sd(p1)
> var(p1)
[,1]
[1,] 100
>
>
> hist(p1, breaks=50, col = rgb(1, 1, 1, 0.5),
+ main = "histogram of p1 and p2",)
> abline(v=mean(p1), col="black", lwd=3)
> hist(p2, add=T, breaks=50, col=rgb(1,1,.5,.5))
> abline(v=mean(p2), col="red", lwd=3)
>
>
> hist(p1, breaks=50, col=rgb(0,.5,.5,.5))
> abline(v=mean(p1),lwd=2)
> abline(v=mean(p1)-sd(p1), lwd=2)
> abline(v=mean(p1)+sd(p1), lwd=2)
> abline(v=c(m.p1-2*sd.p1, m.p1+2*sd.p1), lwd=2, col='red')
> abline(v=c(m.p1-3*sd.p1, m.p1+3*sd.p1), lwd=2, col='green')
>
> # area bet black = 68%
> # between red = 95%
> # between green = 99%
>
> pnorm(m.p1+sd.p1, m.p1, sd.p1)
[1] 0.8413447
> pnorm(m.p1-sd.p1, m.p1, sd.p1)
[1] 0.1586553
> pnorm(m.p1+sd.p1, m.p1, sd.p1) -
+ pnorm(m.p1-sd.p1, m.p1, sd.p1)
[1] 0.6826895
>
> pnorm(m.p1+2*sd.p1, m.p1, sd.p1) -
+ pnorm(m.p1-2*sd.p1, m.p1, sd.p1)
[1] 0.9544997
>
> pnorm(m.p1+3*sd.p1, m.p1, sd.p1) -
+ pnorm(m.p1-3*sd.p1, m.p1, sd.p1)
[1] 0.9973002
>
> m.p1
[1] 100
> sd.p1
[1] 10
>
> # 표준점수 (standardized score)
> za.p1 <- (p1-mean(p1))/sd(p1)
> # e.g.,
> za.eg.1 <- (120 - 100) / 10
> za.eg.1
[1] 2
> # function
> zb.p1 <- scale(p1)
> tail(za.p1 == zb.p1, 10)
[,1]
[999991,] TRUE
[999992,] TRUE
[999993,] TRUE
[999994,] TRUE
[999995,] TRUE
[999996,] TRUE
[999997,] TRUE
[999998,] TRUE
[999999,] TRUE
[1000000,] TRUE
>
> mean(za.p1)
[1] -2.100958e-17
> mean(za.p1)==mean(zb.p1)
[1] TRUE
> round(mean(za.p1),10)
[1] 0
> sd(za.p1)
[1] 1
>
> pnorm(110, 100, 10)
[1] 0.8413447
> pnorm((110-100)/10)
[1] 0.8413447
> pnorm(1)
[1] 0.8413447
> pnorm(90, 100, 10)
[1] 0.1586553
> pnorm((90-100)/10)
[1] 0.1586553
> pnorm(-1)
[1] 0.1586553
> pnorm(110, 100, 10) - pnorm(90, 100, 10)
[1] 0.6826895
> pnorm(1) - pnorm(-1)
[1] 0.6826895
>
> pnorm(120, 100, 10) - pnorm(80, 100, 10)
[1] 0.9544997
> pnorm(2) - pnorm(-2)
[1] 0.9544997
>
>
> pnorm(118, 100, 10)
[1] 0.9640697
> pnorm(82, 100, 10)
[1] 0.03593032
> pnorm(118, 100, 10) - pnorm(82, 100, 10)
[1] 0.9281394
> pnorm(1.8)
[1] 0.9640697
> pnorm(-1.8)
[1] 0.03593032
> pnorm(1.8) - pnorm(-1.8)
[1] 0.9281394
>
> hist(za.p1, breaks=50, col=rgb(1,0,0,0))
> abline(v=c(m.p1, -1.8, 1.8), col='red')
> 1-(pnorm(1.8)-pnorm(-1.8))
[1] 0.07186064
>
> pnorm(1)-pnorm(-1)
[1] 0.6826895
> 1-(pnorm(-1)*2)
[1] 0.6826895
>
> pnorm(2)-pnorm(-2)
[1] 0.9544997
> 1-(pnorm(-2)*2)
[1] 0.9544997
>
> 1-(pnorm(-3)*2)
[1] 0.9973002
>
> #
> hist(p1, breaks=50, col=rgb(.9,.9,.9,.9))
> abline(v=mean(p1),lwd=2)
> abline(v=mean(p1)-sd(p1), lwd=2)
> abline(v=mean(p1)+sd(p1), lwd=2)
> abline(v=c(m.p1-2*sd.p1, m.p1+2*sd.p1), lwd=2, col='red')
> abline(v=c(m.p1-3*sd.p1, m.p1+3*sd.p1), lwd=2, col='green')
>
> # 68%
> a <- qnorm(.32/2)
> b <- qnorm(1-.32/2)
> c(a, b)
[1] -0.9944579 0.9944579
> c(-1, 1)
[1] -1 1
> pnorm(a)
[1] 0.16
> pnorm(b)
[1] 0.84
> pnorm(-1)
[1] 0.1586553
> pnorm(1)
[1] 0.8413447
>
> # 95%
> c <- qnorm(.05/2)
> d <- qnorm(1-.05/2)
> c(c, d)
[1] -1.959964 1.959964
> c(-2,2)
[1] -2 2
>
> # 99%
> e <- qnorm(.01/2)
> f <- qnorm(1-.01/2)
> c(e,f)
[1] -2.575829 2.575829
> c(-3,3)
[1] -3 3
>
> pnorm(b)-pnorm(a)
[1] 0.68
> c(a, b)
[1] -0.9944579 0.9944579
> c(-1, 1)
[1] -1 1
> pnorm(d)-pnorm(c)
[1] 0.95
> c(c, d)
[1] -1.959964 1.959964
> c(-2, 2)
[1] -2 2
> pnorm(f)-pnorm(e)
[1] 0.99
> c(e, f)
[1] -2.575829 2.575829
> c(-3, 3)
[1] -3 3
>
> qnorm(.5)
[1] 0
> qnorm(1)
[1] Inf
>
> ################################
> s.size <- 10
>
> means <- c()
> s1 <- sample(p1, s.size, replace = T)
> mean(s1)
[1] 100.7771
> means <- append(means, mean(s1))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means <- append(means, mean(sample(p1, s.size, replace = T)))
> means
[1] 100.77711 99.34268 101.12569 99.03624 98.73117
> hist(means)
>
> iter <- 100000
> # means <- c()
> means <- rep(NA, iter)
> tail(means)
[1] NA NA NA NA NA NA
>
> for (i in 1:iter) {
+ # means <- append(means, mean(sample(p1, s.size, replace = T)))
+ means[i] <- mean(sample(p1, s.size, replace = T))
+ }
> length(means)
[1] 100000
> mean(means)
[1] 99.99036
> sd(means)
[1] 3.156405
> var(means)
[1] 9.962894
> se.s <- sd(means)
>
> hist(means, breaks=50,
+ xlim = c(mean(means)-5*sd(means), mean(means)+10*sd(means)),
+ col=rgb(1, 1, 1, .5))
> abline(v=mean(means), col="black", lwd=3)
> # now we want to get sd of this distribution
> lo1 <- mean(means)-se.s
> hi1 <- mean(means)+se.s
> lo2 <- mean(means)-2*se.s
> hi2 <- mean(means)+2*se.s
> lo3 <- mean(means)-3*se.s
> hi3 <- mean(means)+3*se.s
> abline(v=mean(means), col="black", lwd=2)
> # abline(v=mean(p2), colo='darkgreen', lwd=2)
> abline(v=c(lo1, hi1, lo2, hi2, lo3, hi3),
+ col=c("red","red", "blue", "blue", "orange", "orange"),
+ lwd=2)
>
> # meanwhile . . . .
> se.s
[1] 3.156405
> sd(means)
[1] 3.156405
> var(means)
[1] 9.962894
>
> se.z <- sqrt(var(p1)/s.size)
> se.z
[,1]
[1,] 3.162278
> var.z <- var(p1)/s.size
> var.z
[,1]
[1,] 10
>
> # sd of sample means (sd(means))
> # = se.s
>
> # when iter value goes to
> # infinite value:
> # mean(means) = mean(p1)
> # and
> # var(means) = var(p1) / s.size
> # sd(means) = sqrt(var(p1)/s.size)
> # that is, se.s = se.z
> # This is called CLT (Central Limit Theorem)
> # see http://commres.net/wiki/cetral_limit_theorem
>
> mean(means)
[1] 99.99036
> var(means)
[1] 9.962894
> sd(means)
[1] 3.156405
> # remember we started talking sample size 10
> mean(p1)
[1] 100
> var(p1)/s.size
[,1]
[1,] 10
> sqrt(var(p1)/s.size)
[,1]
[1,] 3.162278
>
> # because CLT
> loz1 <- mean(p1)-se.z
> hiz1 <- mean(p1)+se.z
> loz2 <- mean(p1)-2*se.z
> hiz2 <- mean(p1)+2*se.z
> loz3 <- mean(p1)-3*se.z
> hiz3 <- mean(p1)+3*se.z
>
> c(lo1, loz1)
[1] 96.83395 96.83772
> c(lo2, loz2)
[1] 93.67755 93.67544
> c(lo3, loz3)
[1] 90.52114 90.51317
>
> c(hi1, hiz1)
[1] 103.1468 103.1623
> c(hi2, hiz2)
[1] 106.3032 106.3246
> c(hi3, hiz3)
[1] 109.4596 109.4868
>
> means.10 <- rnorm2(iter, mean(p1), c(se.z))
> hist(means.10, breaks=50,
+ xlim = c(mean(p1)-5*se.z, mean(p1)+10*se.z),
+ col = rgb(1, 1, 1, .5))
> abline(v=mean(p1), col="black", lwd=3)
> # abline(v=mean(p2), colo='darkgreen', lwd=3)
> abline(v=c(loz1, hiz1, loz2, hiz2, loz3, hiz3),
+ col=c("darkgreen","darkgreen", "blue", "blue", "orange", "orange"),
+ lwd=2)
>
> round(c(lo1, hi1))
[1] 97 103
> round(c(lo2, hi2))
[1] 94 106
> round(c(lo3, hi3))
[1] 91 109
>
> round(c(loz1, hiz1))
[1] 97 103
> round(c(loz2, hiz2))
[1] 94 106
> round(c(loz3, hiz3))
[1] 91 109
>
> m.sample.i.got <- mean(means)+ 1.5*sd(means)
> m.sample.i.got
[1] 104.725
>
> hist(means.10, breaks=30,
+ xlim = c(mean(p1)-7*se.z, mean(p1)+10*se.z),
+ col = rgb(1, 1, 1, .5))
> abline(v=mean(p1), col="black", lwd=3)
> abline(v=m.sample.i.got, col='darkgreen', lwd=3)
>
> # what is the probablity of getting
> # greater than
> # m.sample.i.got?
> m.sample.i.got
[1] 104.725
> pnorm(m.sample.i.got, mean(p1), se.z, lower.tail = F)
[1] 0.06756624
>
> # then, what is the probabilty of getting
> # greater than m.sample.i.got and
> # less than corresponding value, which is
> # mean(means) - m.sample.i.got - mean(means)
> # (green line)
> tmp <- mean(p1) - (m.sample.i.got - mean(p1))
> tmp
[1] 95.27504
> abline(v=tmp, col='red', lwd=3)
> m.sample.i.got
[1] 104.725
> # 붉은 선의 왼쪽 부분을 구해서 2를 곱한 값의 범위
> prob.of.m.sample.i.got <- 2 * pnorm(m.sample.i.got, mean(p1), se.z, lower.tail = F)
> prob.of.m.sample.i.got
[1] 0.1351325
>
> (m.sample.i.got - mean(p1))/se.z
[,1]
[1,] 1.494165
> z.score <- (m.sample.i.got - mean(p1))/se.z
> pnorm(z.score, lower.tail = FALSE)
[,1]
[1,] 0.06756624
> 2 * pnorm(z.score, lower.tail = FALSE)
[,1]
[1,] 0.1351325
>
> # 즉 n = ssize 크기의 샘플을 구했을 때
> # 104.7489 점 오른쪽 그리고,
> # 이에 대응하는 95.25113 지점 왼쪽의
> # 평균이 나오는 probability는
> # 0.1331684
>
> ### one more time
> # this time, with a story
> # 한편 우리는 특정한 평균과 표준편차를 갖는
> # p2를 알고 있다 (110, 10)
> mean(p2)
[1] 110
> sd(p2)
[1] 10
> set.seed(101)
> sample.i.got.p2 <- sample(p2, s.size)
> sample.i.got.p2
[1] 106.63828 107.09795 108.08661 119.09268 103.38813 118.30613 111.49542
[8] 94.25277 115.38045 95.49686
>
> m.sample.i.got.p2 <- mean(sample.i.got.p2)
> m.sample.i.got.p2
[1] 107.9235
>
> # 위의 샘플 평균이 위에서 얘기했던 샘플평균들의 집합에서
> # 나올 확률은 무엇일까?
> hist(means.10, breaks=30,
+ xlim = c(tmp-10*se.z, m.sample.i.got+10*se.z),
+ col = rgb(1, 1, 1, .5))
> abline(v=mean(p1), col="black", lwd=3)
> abline(v=m.sample.i.got.p2, col='blue', lwd=3)
>
> # what is the probablity of getting
> # greater than
> # m.sample.i.got?
> m.sample.i.got.p2
[1] 107.9235
> pnorm(m.sample.i.got.p2, mean(p1), se.z, lower.tail = F)
[1] 0.006111512
> # 혹은
> z.cal <- (m.sample.i.got.p2-mean(p1))/se.z
> pnorm(z.cal, lower.tail = F)
[,1]
[1,] 0.006111512
>
> # then, what is the probability of getting
> # greater than m.sample.i.got and
> # less than corresponding value, which is
> # mean(means) - m.sample.i.got - mean(means)
> # (green line)
> abline(v=mean(p1)-(m.sample.i.got.p2-mean(p1)), col='red', lwd=3)
> 2 * pnorm(m.sample.i.got.p2, mean(p1), se.z, lower.tail = F)
[1] 0.01222302
> 2 * pnorm(z.cal, lower.tail = F)
[,1]
[1,] 0.01222302
>
> # 위는 샘플평균과 모집단 평균값 간의 차이 diff를
> # standard error값으로 즉,
> # sd of sample means (n=s.size) 값으로 나눠 준
> # 표준점수 값을 구하고
> # 이 점수에 대한 probability값을 구해 본것
>
> diff <- m.sample.i.got.p2 - mean(p1)
> se.z <- sqrt(var(p1)/s.size)
> z.cal <- diff / se.z
> z.cal
[,1]
[1,] 2.505639
> prob.z <- pnorm(z.cal, lower.tail=F)*2
> prob.z
[,1]
[1,] 0.01222302
> # 위에 대한 해석 . . . . .
> #
> #
> ###################################################
>
> ###################################################
> # 다른 케이스 --> t-test 케이스
> # what if we do not know the sd of
> # population?
> # we use sd(sample) instead
> # obtained z-score is called t-score
> # and we use t-distribution to get
> # the probability of the t-score instead
> # of normal distribution (z).
> diff
[1] 7.923527
> se.t <- sqrt(var(sample.i.got.p2)/s.size)
> t.cal <- diff/se.t
> t.cal
[1] 2.914599
> pt(t.cal, df=s.size-1, lower.tail = F)
[1] 0.008591086
> pt(t.cal, df=s.size-1, lower.tail = F)*2
[1] 0.01718217
> prob.t <- pt(t.cal, df=s.size-1, lower.tail = F)*2
>
> # note that
> z.cal
[,1]
[1,] 2.505639
> se.z
[,1]
[1,] 3.162278
> prob.z
[,1]
[1,] 0.01222302
>
> t.cal
[1] 2.914599
> se.t
[1] 2.718566
> prob.t
[1] 0.01718217
>
> t.test(sample.i.got.p2, mu=mean(p1))
One Sample t-test
data: sample.i.got.p2
t = 2.9146, df = 9, p-value = 0.01718
alternative hypothesis: true mean is not equal to 100
95 percent confidence interval:
101.7737 114.0733
sample estimates:
mean of x
107.9235
>
> # 그렇다면
> # 두 샘플의 평균과 표준편차만 알고
> # 모집단들의 파라미터는 모를 때는?
>
> # note that
> # v(x)+v(y) = v(x)+v(y)
> # = ss/dfx + ss/dfy
>
> s.size = 30
> s1 <- sample(p1, s.size, replace = T)
> s2 <- sample(p2, s.size, replace = T)
> s1
[1] 100.46855 97.67673 94.48868 97.51996 94.28300 113.78623 95.44617
[8] 92.91108 97.46601 100.66815 113.94452 81.20774 93.77529 93.64388
[15] 111.43702 104.34787 98.82403 100.93909 111.08048 110.12275 94.68502
[22] 100.14734 80.23712 88.02110 118.89602 99.46530 106.92325 97.13274
[29] 106.40479 107.44831
> s2
[1] 120.28360 105.80657 116.28974 106.63910 108.66632 107.51868 123.66436
[8] 96.06523 111.53770 108.78226 115.22628 104.60990 105.07035 114.90022
[15] 103.92860 105.06587 113.77416 108.07776 111.92824 112.01554 104.34549
[22] 100.51113 84.75464 96.36697 117.09381 116.29935 117.20402 109.61657
[29] 106.46650 105.86733
>
> mean(s1)
[1] 100.1133
> mean(s2)
[1] 108.6125
> sd(s1)
[1] 9.132472
> sd(s2)
[1] 7.916677
>
> ss(s1)
[1] 2418.659
> ss(s2)
[1] 1817.54
> df1 <- s.size - 1
> df2 <- s.size - 1
> v.pooled <- (ss(s1)+ss(s2))/(df1+df2)
> se.tst <- sqrt((v.pooled/s.size)+(v.pooled)/s.size)
> diff <- abs(mean(s2)-mean(s1))
> t.ts.cal <- diff/se.tst
> t.ts.cal
[1] 3.851705
> pt(t.ts.cal, df=df1+df2, lower.tail = F)*2
[1] 0.0002954557
>
> t.test(s2, s1, var.equal=T)
Two Sample t-test
data: s2 and s1
t = 3.8517, df = 58, p-value = 0.0002955
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
4.082229 12.916309
sample estimates:
mean of x mean of y
108.6125 100.1133
>
>
> comb <- list(s1 = s1, s2 = s2)
> op <- stack(comb)
> head(op)
values ind
1 100.46855 s1
2 97.67673 s1
3 94.48868 s1
4 97.51996 s1
5 94.28300 s1
6 113.78623 s1
> colnames(op)[1] <- "values"
> colnames(op)[2] <- "group"
> op$group <- factor(op$group)
> head(op)
values group
1 100.46855 s1
2 97.67673 s1
3 94.48868 s1
4 97.51996 s1
5 94.28300 s1
6 113.78623 s1
> boxplot(op$values~op$group)
>
> # Calculate the mean of 'Value' grouped by 'Category'
> m.by.group <- aggregate(values ~ group, data = op, FUN = mean, na.rm = TRUE)
> m.by.group
group values
1 s1 100.1133
2 s2 108.6125
> m.s1 <- m.by.group$values[1]
> m.s2 <- m.by.group$values[2]
> m.tot <- mean(op$values)
> m.s1 <- mean(s1)
> m.s2 <- mean(s2)
>
> n.s1 <- length(s1)
> n.s2 <- length(s2)
> df.s1 <- n.s1-1
> df.s2 <- n.s2-1
>
> ss.tot <- sum((m.tot-op$values)^2)
> ss.tot
[1] 5319.762
> ss.tot <- ss(op$values)
> ss.tot
[1] 5319.762
> ss.bet <- ((m.tot-m.s1)^2*n.s1) +
+ ((m.tot-m.s2)^2*n.s2)
> ss.wit <- ss(s1) + ss(s2)
> ss.tot
[1] 5319.762
> ss.bet
[1] 1083.564
> ss.wit
[1] 4236.199
> ss.bet+ss.wit
[1] 5319.762
>
> df.tot <- (n.s1+n.s2)-1
> df.bet <- nlevels(op$group)-1
> df.wit <- df.s1+df.s2
>
> ms.tot <- ss.tot / df.tot
> ms.bet <- ss.bet / df.bet
> ms.wit <- ss.wit / df.wit
>
> r.sq <- ss.bet / ss.tot
> r.sq
[1] 0.2036865
> f.cal <- ms.bet / ms.wit
> f.cal
[1] 14.83563
> prob.f <- pf(f.cal, df.bet, df.wit, lower.tail = F)
> prob.f
[1] 0.0002954557
>
> ss.bet
[1] 1083.564
> df.bet
[1] 1
> ms.bet
[1] 1083.564
> ss.wit
[1] 4236.199
> df.wit
[1] 58
> ms.wit
[1] 73.03791
> f.cal
[1] 14.83563
> prob.f
[1] 0.0002954557
>
> m.f <- aov(values~group, data=op)
> summary(m.f)
Df Sum Sq Mean Sq F value Pr(>F)
group 1 1084 1084 14.84 0.000295 ***
Residuals 58 4236 73
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
>
> m.t <- t.test(s2, s1, var.equal = T)
> m.t
Two Sample t-test
data: s2 and s1
t = 3.8517, df = 58, p-value = 0.0002955
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
4.082229 12.916309
sample estimates:
mean of x mean of y
108.6125 100.1133
> m.t$statistic^2
t
14.83563
> m.t$p.value
[1] 0.0002954557
>
> m.l <- lm(values~group, data=op)
> summary(m.l)
Call:
lm(formula = values ~ group, data = op)
Residuals:
Min 1Q Median 3Q Max
-23.8579 -4.3671 -0.5914 6.3721 18.7827
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 100.113 1.560 64.162 < 2e-16 ***
groups2 8.499 2.207 3.852 0.000295 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 8.546 on 58 degrees of freedom
Multiple R-squared: 0.2037, Adjusted R-squared: 0.19
F-statistic: 14.84 on 1 and 58 DF, p-value: 0.0002955
> f.cal
[1] 14.83563
> df.bet
[1] 1
> df.wit
[1] 58
> prob.f
[1] 0.0002954557
> r.sq
[1] 0.2036865
> anova(m.l)
Analysis of Variance Table
Response: values
Df Sum Sq Mean Sq F value Pr(>F)
group 1 1083.6 1083.56 14.836 0.0002955 ***
Residuals 58 4236.2 73.04
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
> ss.bet
[1] 1083.564
> ss.wit
[1] 4236.199
> ms.bet
[1] 1083.564
> ms.wit
[1] 73.03791
> f.cal
[1] 14.83563
> prob.f
[1] 0.0002954557
>
exercise.1
위에서 p2의 parameter에 대해서 잘 모른다는 점에 주목하라. 그리고 아래 시나리오를 상상하라.
- 어느 한 모집단의 IQ 평균이 100 이고 표준편차가 10 임을 알고 있다. 확률과통계 교수는 머리가 좋아지는 약을 개발하여 이를 팔아보려고 하고 있다. 이를 위해서 확통교수는 25명의 학생에게 머리가 좋아지는 약을 복용하도록 한 후에 IQ를 측정하였다. 그런데, 그 IQ 평균이 106.45 이다. 이 점수를 가지고 약의 효과가 있는지 검증을 해보력고 한다.
- 똑 같은 경우이지만 모집단의 평균을 100으로 추정하고 있지 표준편차는 모르는 상태이다. 그런데, 25명에게 약을 복용시키고 IQ를 측정하니 점수가 105.50이 나왔고, 표준편차는 9.4였다. 이 점수로 약의 효과가 있었는지 검증을 하려 한다.