This is an old revision of the document!
Table of Contents
Recap
Distribution of Sample Means – mu = 40, sigma = 4 (hence var = 16) 인 모집단에서 n = n 사이즈의 샘플링을 무한 반복할 때 그 샘플평균들이 모인 집합
rscript01
rm(list=ls())
rnorm2 <- function(n,mean,sd){
mean+sd*scale(rnorm(n))
}
ss <- function(x) {
sum((x-mean(x))^2)
}
mu <- 40
sigma <- 4
iter <- 10000000
sz <- 16
se <- sigma/sqrt(sz)
################################
means <- rnorm2(iter, mu, se)
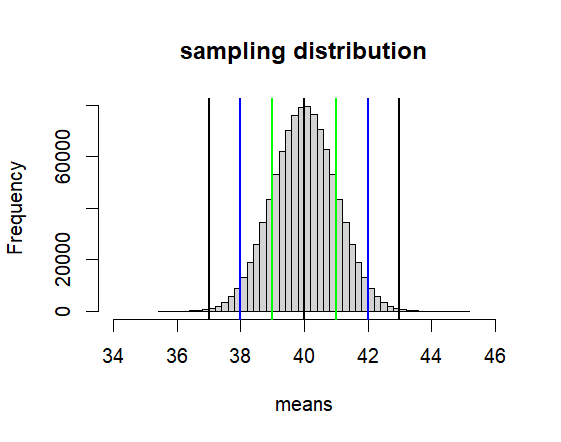
hist(means, breaks=50,
xlim = c(mu-6*se, mu+6*se),
main = paste("sampling distribution"))
abline(v=mu, col='black', lwd=2)
one <- -1
two <- -2
thr <- -3
lo1 <- mu + se*one
hi1 <- mu - se*one
lo2 <- mu + se*two
hi2 <- mu - se*two
lo3 <- mu + se*thr
hi3 <- mu - se*thr
abline(v=c(lo1, lo2, lo3, hi1, hi2, hi3),
col=c("green","blue", "black"),
lwd=2)
print(c(lo1, hi1))
print(c(lo2, hi2))
print(c(lo3, hi3))
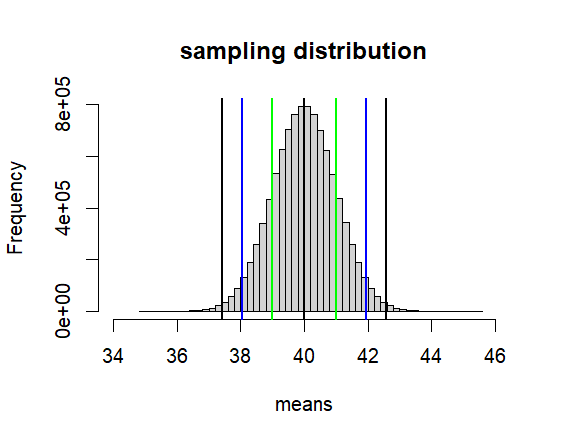
hist(means, breaks=50,
xlim = c(mu-6*se, mu+6*se),
main = paste("sampling distribution"))
abline(v=mu, col='black', lwd=2)
one <- qnorm(.32/2)
two <- qnorm(.05/2)
thr <- qnorm(.01/2)
lo1 <- mu - se*one
hi1 <- mu + se*one
lo2 <- mu - se*two
hi2 <- mu + se*two
lo3 <- mu - se*thr
hi3 <- mu + se*thr
abline(v=c(lo1, lo2, lo3, hi1, hi2, hi3),
col=c("green","blue", "black"),
lwd=2)
c(lo1, hi1)
c(lo2, hi2)
c(lo3, hi3)
# 위를 이해하고 나면
# 어느 한 샘플의 (사이즈는 n이어야 한ㄷ)
# 평균이 위의 모집단에서 나올 확률을
# 알아 볼 수 있다.
m.samp <- 37
p.val <- pnorm(m.samp, mu, se)*2
p.val
z.cal <- (m.samp-mu)/se
z.cal
p.val <- pnorm(z.cal)*2
p.val
zmeans <- scale(means)
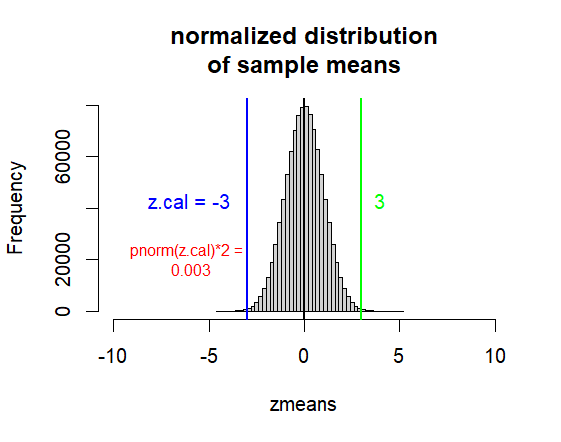
hist(zmeans, breaks=50,
xlim = c(0-10*1, 0+10*1),
main=("normalized distribution\nof sample means"))
abline(v=0, col="black", lwd=2)
abline(v=z.cal, col='blue', lwd=2)
abline(v=-z.cal, col="green", lwd=2)
text(x=-6, y=50000,
label=paste("z.cal =", z.cal),
pos = 1,
col="blue", cex=1)
text(x=4, y=50000,
label=paste(-z.cal),
pos=1,
col="green", cex=1)
text(x=-6, y=30000,
label=paste("pnorm(z.cal)*2 =", "\n",
round(p.val,3)),
pos = 1,
col="red", cex=.8)
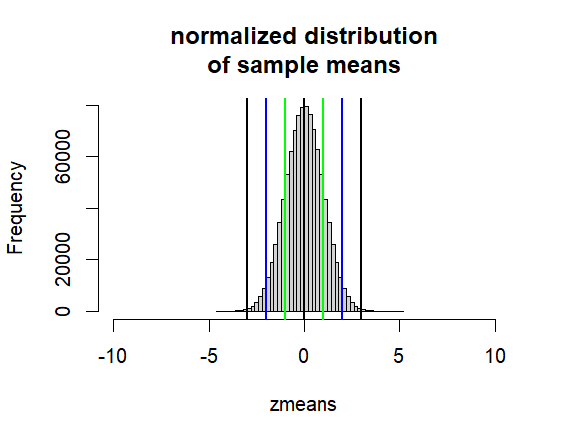
hist(zmeans, breaks=50,
xlim = c(0-10*1, 0+10*1),
main=("normalized distribution\nof sample means"))
abline(v=0, col="black", lwd=2)
abline(v=c(-1,-2,-3,1,2,3),
col=c("green", "blue", "black"), lwd=2)
z.cal
p.val
#####
# 위의 아이디어로는 z.cal 점수가
# +-2 밖에 있는지 보면 된다. 즉,
# 이는 prob가 0.05보다 작은지
# 보면 되는 것이다.
#####
# +-2 는 정확한 숫자가 아니고
# qnorm(.05/2) 에 해당하는 숫자
# 가 정확한 숫자
two.minus.exact <- qnorm(.05/2)
two.plus.exact <- qnorm(1-(.05/2))
c(two.minus.exact, two.plus.exact)
#####
# 그러나 R 사용시에는 z 점수로
# 판단하기 보다는
# 직접 구하는 prob.로 판단
pnorm(z.cal)*2
p.val
#####
# 위에서 그룹 간의 차이를
# standard error로 나누는 것에 주의
#
################
m.samp <- 43
sd.samp <- 4
sz <- 16
samp <- rnorm2(sz, m.samp, sd.samp)
diff <- m.samp - mu
se <- sd.samp / sqrt(sz)
t.cal <- diff/se
df <- sz-1
p.val <- pt(t.cal, df=df, lower.tail = F)*2
t.cal
df
p.val
t.test(samp, mu=mu)
out01
>
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
> mu <- 40
> sigma <- 4
> iter <- 10000000
> sz <- 16
> se <- sigma/sqrt(sz)
> ################################
> means <- rnorm2(iter, mu, se)
> hist(means, breaks=50,
+ xlim = c(mu-6*se, mu+6*se),
+ main = paste("sampling distribution"))
> abline(v=mu, col='black', lwd=2)
>
> one <- -1
> two <- -2
> thr <- -3
>
> lo1 <- mu + se*one
> hi1 <- mu - se*one
> lo2 <- mu + se*two
> hi2 <- mu - se*two
> lo3 <- mu + se*thr
> hi3 <- mu - se*thr
>
> abline(v=c(lo1, lo2, lo3, hi1, hi2, hi3),
+ col=c("green","blue", "black"),
+ lwd=2)
> print(c(lo1, hi1))
[1] 39 41
> print(c(lo2, hi2))
[1] 38 42
> print(c(lo3, hi3))
[1] 37 43
>
> hist(means, breaks=50,
+ xlim = c(mu-6*se, mu+6*se),
+ main = paste("sampling distribution"))
> abline(v=mu, col='black', lwd=2)
>
> one <- qnorm(.32/2)
> two <- qnorm(.05/2)
> thr <- qnorm(.01/2)
>
> lo1 <- mu - se*one
> hi1 <- mu + se*one
> lo2 <- mu - se*two
> hi2 <- mu + se*two
> lo3 <- mu - se*thr
> hi3 <- mu + se*thr
>
> abline(v=c(lo1, lo2, lo3, hi1, hi2, hi3),
+ col=c("green","blue", "black"),
+ lwd=2)
> c(lo1, hi1)
[1] 40.99446 39.00554
> c(lo2, hi2)
[1] 41.95996 38.04004
> c(lo3, hi3)
[1] 42.57583 37.42417
>
> # 위를 이해하고 나면
> # 어느 한 샘플의 (사이즈는 n이어야 한ㄷ)
> # 평균이 위의 모집단에서 나올 확률을
> # 알아 볼 수 있다.
>
>
> m.samp <- 37
> p.val <- pnorm(m.samp, mu, se)*2
> p.val
[1] 0.002699796
> z.cal <- (m.samp-mu)/se
> z.cal
[1] -3
> p.val <- pnorm(z.cal)*2
> p.val
[1] 0.002699796
>
> zmeans <- scale(means)
> hist(zmeans, breaks=50,
+ xlim = c(0-10*1, 0+10*1),
+ main=("normalized distribution\nof sample means"))
> abline(v=0, col="black", lwd=2)
> abline(v=z.cal, col='blue', lwd=2)
> abline(v=-z.cal, col="green", lwd=2)
> text(x=-6, y=50000,
+ label=paste("z.cal =", z.cal),
+ pos = 1,
+ col="blue", cex=1)
> text(x=4, y=50000,
+ label=paste(-z.cal),
+ pos=1,
+ col="green", cex=1)
> text(x=-6, y=30000,
+ label=paste("pnorm(z.cal)*2 =", "\n",
+ round(p.val,3)),
+ pos = 1,
+ col="red", cex=.8)
>
> hist(zmeans, breaks=50,
+ xlim = c(0-10*1, 0+10*1),
+ main=("normalized distribution\nof sample means"))
> abline(v=0, col="black", lwd=2)
> abline(v=c(-1,-2,-3,1,2,3),
+ col=c("green", "blue", "black"), lwd=2)
>
> z.cal
[1] -3
> p.val
[1] 0.002699796
> #####
> # 위의 아이디어로는 z.cal 점수가
> # +-2 밖에 있는지 보면 된다. 즉,
> # 이는 prob가 0.05보다 작은지
> # 보면 되는 것이다.
> #####
> # +-2 는 정확한 숫자가 아니고
> # qnorm(.05/2) 에 해당하는 숫자
> # 가 정확한 숫자
> two.minus.exact <- qnorm(.05/2)
> two.plus.exact <- qnorm(1-(.05/2))
> c(two.minus.exact, two.plus.exact)
[1] -1.959964 1.959964
> #####
> # 그러나 R 사용시에는 z 점수로
> # 판단하기 보다는
> # 직접 구하는 prob.로 판단
> pnorm(z.cal)*2
[1] 0.002699796
> p.val
[1] 0.002699796
> #####
> # 위에서 그룹 간의 차이를
> # standard error로 나누는 것에 주의
> #
>
>
> ################
> m.samp <- 43
> sd.samp <- 4
> sz <- 16
> samp <- rnorm2(sz, m.samp, sd.samp)
> diff <- m.samp - mu
> se <- sd.samp / sqrt(sz)
> t.cal <- diff/se
> df <- sz-1
> p.val <- pt(t.cal, df=df, lower.tail = F)*2
> t.cal
[1] 3
> df
[1] 15
> p.val
[1] 0.008972737
> t.test(samp, mu=mu)
One Sample t-test
data: samp
t = 3, df = 15, p-value = 0.008973
alternative hypothesis: true mean is not equal to 40
95 percent confidence interval:
40.86855 45.13145
sample estimates:
mean of x
43
>
rscript02
##### # m.a <- 5.8 m.b <- 6.3 sd.a <- .5 sd.b <- .5 sz.a <- 16 sz.b <- 16 df.a <- sz.a-1 df.b <- sz.b-1 df <- df.a + df.b a <- rnorm2(sz.a, m.a, sd.a) b <- rnorm2(sz.b, m.b, sd.b) diff <- m.a - m.b pv <- (ss(a)+ss(b))/(df.a+df.b) se <- sqrt(pv/sz.a+pv/sz.b) t.cal <- diff / se p.val <- pt(t.cal, df=df)*2 diff se t.cal df p.val t.test(a,b, var.equal = T) diff - se*2 diff + se*2 lo <- qt(.05/2,df) lo hi <- -lo diff + se*lo diff + se*hi ##### # t-test repeated measre ##### m.t1 <- 103 m.t2 <- 111 sd.t1 <- 10 sd.t2 <- 10 sz <- 16 t1 <- rnorm2(sz, m.t1, sd.t1) t2 <- rnorm2(sz, m.t2, sd.t2) t1 t2 mdiff <- m.t1-m.t2 diff <- t1-t2 sd.diff <- sd(diff) se <- sd.diff/sqrt(sz) t.cal <- mdiff/se p.val <- pt(t.cal, df=sz-1)*2 t.cal sz-1 p.val t.test(t1,t2, paired=T) two <- qt(.05/2, df=sz-1) two lo <- se*two hi <- -lo c(lo, hi) c(mdiff+lo, mdiff+hi)
rout02
> #####
> #
> m.a <- 5.8
> m.b <- 6.3
> sd.a <- .5
> sd.b <- .5
> sz.a <- 16
> sz.b <- 16
> df.a <- sz.a-1
> df.b <- sz.b-1
> df <- df.a + df.b
> a <- rnorm2(sz.a, m.a, sd.a)
> b <- rnorm2(sz.b, m.b, sd.b)
> diff <- m.a - m.b
> pv <- (ss(a)+ss(b))/(df.a+df.b)
> se <- sqrt(pv/sz.a+pv/sz.b)
> t.cal <- diff / se
> p.val <- pt(t.cal, df=df)*2
>
> diff
[1] -0.5
> se
[1] 0.1767767
> t.cal
[1] -2.828427
> df
[1] 30
> p.val
[1] 0.008257336
> t.test(a,b, var.equal = T)
Two Sample t-test
data: a and b
t = -2.8284, df = 30, p-value = 0.008257
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-0.8610262 -0.1389738
sample estimates:
mean of x mean of y
5.8 6.3
> diff - se*2
[1] -0.8535534
> diff + se*2
[1] -0.1464466
> lo <- qt(.05/2,df)
> lo
[1] -2.042272
> hi <- -lo
> diff + se*lo
[1] -0.8610262
> diff + se*hi
[1] -0.1389738
>
> #####
> # t-test repeated measre
> #####
> m.t1 <- 103
> m.t2 <- 111
> sd.t1 <- 10
> sd.t2 <- 10
> sz <- 16
> t1 <- rnorm2(sz, m.t1, sd.t1)
> t2 <- rnorm2(sz, m.t2, sd.t2)
> t1
[,1]
[1,] 89.58295
[2,] 97.20986
[3,] 100.82700
[4,] 120.11867
[5,] 103.06410
[6,] 117.36762
[7,] 98.82191
[8,] 111.72472
[9,] 100.06093
[10,] 114.58757
[11,] 105.99472
[12,] 84.34803
[13,] 94.63867
[14,] 94.49667
[15,] 106.03514
[16,] 109.12144
attr(,"scaled:center")
[1] 0.08912759
attr(,"scaled:scale")
[1] 0.9759765
> t2
[,1]
[1,] 114.76609
[2,] 111.81937
[3,] 102.93248
[4,] 122.85959
[5,] 105.68180
[6,] 110.43890
[7,] 115.34844
[8,] 97.39180
[9,] 117.00475
[10,] 98.63924
[11,] 118.87807
[12,] 107.55519
[13,] 128.46569
[14,] 93.50094
[15,] 107.15280
[16,] 123.56487
attr(,"scaled:center")
[1] 0.2000755
attr(,"scaled:scale")
[1] 0.8946962
> mdiff <- m.t1-m.t2
> diff <- t1-t2
> sd.diff <- sd(diff)
> se <- sd.diff/sqrt(sz)
> t.cal <- mdiff/se
> p.val <- pt(t.cal, df=sz-1)*2
> t.cal
[1] -2.2741
> sz-1
[1] 15
> p.val
[1] 0.03808083
> t.test(t1,t2, paired=T)
Paired t-test
data: t1 and t2
t = -2.2741, df = 15, p-value = 0.03808
alternative hypothesis: true mean difference is not equal to 0
95 percent confidence interval:
-15.4981736 -0.5018264
sample estimates:
mean difference
-8
> two <- qt(.05/2, df=sz-1)
> two
[1] -2.13145
> lo <- se*two
> hi <- -lo
> c(lo, hi)
[1] -7.498174 7.498174
> c(mdiff+lo, mdiff+hi)
[1] -15.4981736 -0.5018264
>
>
Sum up
see t-test
- 샘플사이즈가 크면 t-test는 z-test와 같은 결과를 나타나기에 대부분 t-test를 사용함.
- t-test는 기본적으로 두 집합의 (그룹의) 평균의 차이를 판단하는 방법
- 자세한 상황으로 4가지가 있음
- mu, sigma를 모두 알고 있는 상태에서 한 그룹의 평균을 mu와 비교해서 차이를 알고 싶을 때 – z-test를 할 수 있는 상황
- mu만 알고 있고, 그룹의 평균과 표준편차를 알고 있고, mu와 그룹평균의 차이를 알고 싶을 때 – t-test (one sample)
- 모집단의 파라미터는 모르고, 두 그룹의 평균과 표준편차만을 알고 있어서, 이 두 그룹의 평균의 차이를 알고 싶을 때 – t-test (two sample, two sample independent, independent samples t-test)
- 모집단의 파라미터는 모르는 상태에서, 한 그룹이 반복 측정되었을 때 – t-test (repeated measure t-test, paired sample t-test)
- 각 사정에 따라서 standard error값을 구하는 방법이 달라짐
- t-test는 그룹 간의 차이를 (R에서 diff 변인으로 예를 든) – 독립변인 때문에 생긴 효과 (차이)
- se 값으로 나눠줘서 – 랜덤차이 (표준오차 = standard error) – a sample mean's random error from the mean
- 샘플의 평균이 랜덤하게 모집단의 평균에서 얼마나 떨어져서 나타나는 가를 나타내주는 단위 = standard error
\begin{eqnarray}
z\;\;\;\text{or}\;\;\;t & = & \frac{\overline{X}-\mu}{\sigma_{\overline{X}} }, \quad \text{where } \;\; \sigma_{\overline{X}} = \frac{\sigma}{\sqrt{n}} \\
t & = & \frac{ \overline{X}-\mu}{s_{\overline{X}} }, \quad \text{where } \;\; s_{\overline{X}} = \frac{s}{\sqrt{n}} \\
t & = & \frac{(\overline{X_a}-\overline{X_b})-(\mu_a-\mu_b)}{\sigma_{\text{diff} }} , \;\;\; \\
& & \qquad \qquad \;\;\; \text{where } \;\; \sigma_{\text{diff} } = \sqrt{ \frac{s^2_{\text{pooled}}}{n_a} + \frac{s^2_{\text{pooled}}}{n_b} } \nonumber \\
& & \qquad \qquad \;\;\; s^2_{\text{pooled}} = \frac {\text{SS}_a + \text{SS}_b} {df_a + df_b} \nonumber \\
t & = & \frac{\overline{D}-0}{s_{ \overline{D} }}, \quad \text{where } \;\; s_{ \overline{D} } = \frac {s_D}{\sqrt{n}} \\
\end{eqnarray}
위의 4 경우의 공통점은 :
\begin{eqnarray*}
t & = & \displaystyle \frac{\text{obtained difference}}{\text{difference by random error}}
\end{eqnarray*}
Two sample t-test
NOT convinced?
\begin{eqnarray*} E \left[ \overline{X} - \overline{Y} \right] & = & E \left[ \overline{X} \right] - E \left[ \overline{Y} \right] \\ V \left[ \overline{X} - \overline{Y} \right] & = & V \left[ \overline{X} \right] + V \left[ \overline{Y} \right] \\ & = & \frac {\sigma_{X}} {n_{X}} + \frac {\sigma_{Y}} {n_{Y}} \\ \end{eqnarray*}
rs.two.sample.t.test
rm(list=ls())
rnorm2 <- function(n,mean,sd){
mean+sd*scale(rnorm(n))
}
ss <- function(x) {
sum((x-mean(x))^2)
}
N.p <- 1000000
m.p <- 100
sd.p <- 10
set.seed(82)
p1 <- rnorm2(N.p, m.p, sd.p)
mean(p1)
sd(p1)
p2 <- rnorm2(N.p, m.p+10, sd.p)
mean(p2)
sd(p2)
s.size <- 25
iter <- 100000
# means <- c()
mdiffs <- rep(NA, iter)
means.s1 <- rep(NA, iter)
means.s2 <- rep(NA, iter)
tail(mdiffs)
for (i in 1:iter) {
# means <- append(means, mean(sample(p1, s.size, replace = T)))
s1 <- sample(p1, s.size, replace = T)
s2 <- sample(p2, s.size, replace = T)
means.s1[i] <- mean(s1)
means.s2[i] <- mean(s2)
mdiffs[i] <- mean(s2-s1)
}
head(mdiffs)
tail(mdiffs)
m.diff <- mean(mdiffs)
var.diff <- var(mdiffs)
sd.diff <- sd(mdiffs)
m.diff
var.diff
sd.diff
var(means.s1)
var(p1)/s.size
var(means.s2)
var(p2)/s.size
var(means.s1-means.s2)
var(means.s1) + var(means.s2)
var(p1)/s.size + var(p2)/s.size
var.diff <- (var(p1)/s.size) + (var(p2)/s.size)
var.diff
sqrt(var.diff)
se.diff <- sqrt(var.diff)
se.diff
s1 <- sample(p1, s.size, replace = T)
s2 <- sample(p2, s.size, replace = T)
df <- s.size - 1
pv <- (ss(s1)+ss(s2))/(df+df)
pv
ms1 <- ss(s1)/df
ms2 <- ss(s2)/df
ms1
ms2
(ms1+ms2)/2
se <- sqrt(ms1/s.size+ms2/s.size)
se
se.z <- sqrt(pv/s.size+pv/s.size)
se.z
diff <- mean(s2)-mean(s1)
t.cal <- diff / se.z
t.test(s1,s2, var.equal = T)
t.cal
pt(abs(t.cal), df+df, lower.tail = F)*2
df+df
ro.two.sample.t.test
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
>
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
> N.p <- 1000000
> m.p <- 100
> sd.p <- 10
>
> set.seed(82)
> p1 <- rnorm2(N.p, m.p, sd.p)
> mean(p1)
[1] 100
> sd(p1)
[1] 10
>
> p2 <- rnorm2(N.p, m.p+10, sd.p)
> mean(p2)
[1] 110
> sd(p2)
[1] 10
>
> s.size <- 25
>
> iter <- 100000
> # means <- c()
> mdiffs <- rep(NA, iter)
> means.s1 <- rep(NA, iter)
> means.s2 <- rep(NA, iter)
> tail(mdiffs)
[1] NA NA NA NA NA NA
>
> for (i in 1:iter) {
+ # means <- append(means, mean(sample(p1, s.size, replace = T)))
+ s1 <- sample(p1, s.size, replace = T)
+ s2 <- sample(p2, s.size, replace = T)
+ means.s1[i] <- mean(s1)
+ means.s2[i] <- mean(s2)
+ mdiffs[i] <- mean(s2-s1)
+ }
> head(mdiffs)
[1] 8.676338 9.405720 10.395752 10.939396 9.912317 9.549864
> tail(mdiffs)
[1] 13.118479 13.736393 12.947016 9.947961 6.951695 4.664188
> m.diff <- mean(mdiffs)
> var.diff <- var(mdiffs)
> sd.diff <- sd(mdiffs)
> m.diff
[1] 10.00197
> var.diff
[1] 8.023544
> sd.diff
[1] 2.832586
>
> var(means.s1)
[1] 4.007843
> var(p1)/s.size
[,1]
[1,] 4
> var(means.s2)
[1] 4.009957
> var(p2)/s.size
[,1]
[1,] 4
> var(means.s1-means.s2)
[1] 8.023544
> var(means.s1) + var(means.s2)
[1] 8.017801
> var(p1)/s.size + var(p2)/s.size
[,1]
[1,] 8
> var.diff <- (var(p1)/s.size) + (var(p2)/s.size)
> var.diff
[,1]
[1,] 8
> sqrt(var.diff)
[,1]
[1,] 2.828427
> se.diff <- sqrt(var.diff)
> se.diff
[,1]
[1,] 2.828427
>
> s1 <- sample(p1, s.size, replace = T)
> s2 <- sample(p2, s.size, replace = T)
>
> df <- s.size - 1
> pv <- (ss(s1)+ss(s2))/(df+df)
> pv
[1] 84.76206
> ms1 <- ss(s1)/df
> ms2 <- ss(s2)/df
> ms1
[1] 102.4864
> ms2
[1] 67.03775
> (ms1+ms2)/2
[1] 84.76206
>
> se <- sqrt(ms1/s.size+ms2/s.size)
> se
[1] 2.604029
> se.z <- sqrt(pv/s.size+pv/s.size)
> se.z
[1] 2.604029
>
> diff <- mean(s2)-mean(s1)
> t.cal <- diff / se.z
>
> t.test(s1,s2, var.equal = T)
Two Sample t-test
data: s1 and s2
t = -2.9772, df = 48, p-value = 0.004548
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-12.988542 -2.517041
sample estimates:
mean of x mean of y
102.4374 110.1902
> t.cal
[1] 2.97723
> pt(abs(t.cal), df+df, lower.tail = F)*2
[1] 0.004547918
> df+df
[