| BOS | NY | DC | MIA | CHI | SEA | SF | LA | DEN | |
| BOS | 0 | 206 | 429 | 1504 | 963 | 2976 | 3095 | 2979 | 1949 |
| NY | 206 | 0 | 233 | 1308 | 802 | 2815 | 2934 | 2786 | 1771 |
| DC | 429 | 233 | 0 | 1075 | 671 | 2684 | 2799 | 2631 | 1616 |
| MIA | 1504 | 1308 | 1075 | 0 | 1329 | 3273 | 3053 | 2687 | 2037 |
| CHI | 963 | 802 | 671 | 1329 | 0 | 2013 | 2142 | 2054 | 996 |
| SEA | 2976 | 2815 | 2684 | 3273 | 2013 | 0 | 808 | 1131 | 1307 |
| SF | 3095 | 2934 | 2799 | 3053 | 2142 | 808 | 0 | 379 | 1235 |
| LA | 2979 | 2786 | 2631 | 2687 | 2054 | 1131 | 379 | 0 | 1059 |
| DEN | 1949 | 1771 | 1616 | 2037 | 996 | 1307 | 1235 | 1059 | 0 |
- Start by assigning each item to its own cluster, so that if you have N items, you now have N clusters, each containing just one item. Let the distances (similarities) between the clusters equal the distances (similarities) between the items they contain.
- Find the closest (most similar) pair of clusters and merge them into a single cluster, so that now you have one less cluster.
- Compute distances (similarities) between the new cluster and each of the old clusters.
- Repeat steps 2 and 3 until all items are clustered into a single cluster of size N.
Step 3 can be done in different ways, which is what distinguishes single-link from complete-link and average-link clustering.
- single-link clustering (also called the connectedness or minimum method) = the shortest distance from any member of one cluster to any member of the other cluster.
- complete-link clustering (also called the diameter or maximum method) = the longest distance from any member of one cluster to any member of the other cluster.
- average-link clustering = the average distance from any member of one cluster to any member of the other cluster.
| BOS | NY | DC | MIA | CHI | SEA | SF | LA | DEN | |
| BOS | 0 | 206 | 429 | 1504 | 963 | 2976 | 3095 | 2979 | 1949 |
| NY | 206 | 0 | 233 | 1308 | 802 | 2815 | 2934 | 2786 | 1771 |
| DC | 429 | 233 | 0 | 1075 | 671 | 2684 | 2799 | 2631 | 1616 |
| MIA | 1504 | 1308 | 1075 | 0 | 1329 | 3273 | 3053 | 2687 | 2037 |
| CHI | 963 | 802 | 671 | 1329 | 0 | 2013 | 2142 | 2054 | 996 |
| SEA | 2976 | 2815 | 2684 | 3273 | 2013 | 0 | 808 | 1131 | 1307 |
| SF | 3095 | 2934 | 2799 | 3053 | 2142 | 808 | 0 | 379 | 1235 |
| LA | 2979 | 2786 | 2631 | 2687 | 2054 | 1131 | 379 | 0 | 1059 |
| DEN | 1949 | 1771 | 1616 | 2037 | 996 | 1307 | 1235 | 1059 | 0 |
- 가장 가까운 거리의 도시: BOS 와 NY, 206
- 두 도시를 합하여 BOS/NY로 하고 다시 이를 포함한 도시들 간의 거리를 구함
- single link 방법을 사용한다면 BOS/NY와 DC간의 거리는 223이 됨 (가장 가까운 거리를 클러스터와의 거리로 환산하는 방법이 single link method). 마찬가지로 DEN까지의 거리는 1771이 됨
| BOS/NY | DC | MIA | CHI | SEA | SF | LA | DEN | |
| BOS/NY | 0 | 223 | 1308 | 802 | 2815 | 2934 | 2786 | 1771 |
| DC | 223 | 0 | 1075 | 671 | 2684 | 2799 | 2631 | 1616 |
| MIA | 1308 | 1075 | 0 | 1329 | 3273 | 3053 | 2687 | 2037 |
| CHI | 802 | 671 | 1329 | 0 | 2013 | 2142 | 2054 | 996 |
| SEA | 2815 | 2684 | 3273 | 2013 | 0 | 808 | 1131 | 1307 |
| SF | 2934 | 2799 | 3053 | 2142 | 808 | 0 | 379 | 1235 |
| LA | 2786 | 2631 | 2687 | 2054 | 1131 | 379 | 0 | 1059 |
| DEN | 1771 | 1616 | 2037 | 996 | 1307 | 1235 | 1059 | 0 |
- BOS/NY와 가장 가까운 거리의 도시는 DC이고 거리는 223
- BOS/NY/DC 로 클러스터링하고 이와 다른 도시들, 그리고 각 도시들 간의 거리를 다시 계산
| BOS/NY/DC | MIA | CHI | SEA | SF | LA | DEN | |
| BOS/NY/DC | 0 | 1075 | 671 | 2684 | 2799 | 2631 | 1616 |
| MIA | 1075 | 0 | 1329 | 3273 | 3053 | 2687 | 2037 |
| CHI | 671 | 1329 | 0 | 2013 | 2142 | 2054 | 996 |
| SEA | 2684 | 3273 | 2013 | 0 | 808 | 1131 | 1307 |
| SF | 2799 | 3053 | 2142 | 808 | 0 | 379 | 1235 |
| LA | 2631 | 2687 | 2054 | 1131 | 379 | 0 | 1059 |
| DEN | 1616 | 2037 | 996 | 1307 | 1235 | 1059 | 0 |
- 위에서 가장 가까운 도시들 간의 거리는 379이고 이는 SF와 LA 간의 거리
- SF/LA로 합치고 다시 계산하여 매트릭스를 구함
| BOS/NY/DC | MIA | CHI | SEA | SF/LA | DEN | |
| BOS/NY/DC | 0 | 1075 | 671 | 2684 | 2631 | 1616 |
| MIA | 1075 | 0 | 1329 | 3273 | 2687 | 2037 |
| CHI | 671 | 1329 | 0 | 2013 | 2054 | 996 |
| SEA | 2684 | 3273 | 2013 | 0 | 808 | 1307 |
| SF/LA | 2631 | 2687 | 2054 | 808 | 0 | 1059 |
| DEN | 1616 | 2037 | 996 | 1307 | 1059 | 0 |
- 이제 CHI가 BOS/NY/DC/CHI와 가장 가까움 (671)
- BOS/NY/DC/CHI로 병합
| BOS/NY/ DC/CHI | MIA | SEA | SF/LA | DEN | |
| BOS/NY/ DC/CHI | 0 | 1075 | 2013 | 2054 | 996 |
| MIA | 1075 | 0 | 3273 | 2687 | 2037 |
| SEA | 2013 | 3273 | 0 | 808 | 1307 |
| SF/LA | 2054 | 2687 | 808 | 0 | 1059 |
| DEN | 996 | 2037 | 1307 | 1059 | 0 |
- 같은 방법으로 SEA을 SF/LA에 병합 (SF/LA/SEA)
| BOS/NY/ DC/CHI | MIA | SF/LA /SEA | DEN | |
| BOS/NY/ DC/CHI | 0 | 1075 | 2013 | 996 |
| MIA | 1075 | 0 | 2687 | 2037 |
| SF/LA/ SEA | 2054 | 2687 | 0 | 1059 |
| DEN | 996 | 2037 | 1059 | 0 |
| BOS/NY/DC/ CHI/DEN | MIA | SF/LA/SEA | |
| BOS/NY/DC/ CHI/DEN | 0 | 1075 | 1059 |
| MIA | 1075 | 0 | 2687 |
| SF/LA/SEA | 1059 | 2687 | 0 |
| BOS/NY/DC/CHI/ DEN/SF/LA/SEA | MIA | |
| BOS/NY/DC/CHI/ DEN/SF/LA/SEA | 0 | 1075 |
| MIA | 1075 | 0 |
JOHNSON'S HIERARCHICAL CLUSTERING
--------------------------------------------------------------------------------
Method: SINGLE_LINK (minimum distance)
Type of Data: Dissimilarities
Input dataset: cities (D:\Users\Hyo\Documents\UCINET data\Cities\cities)
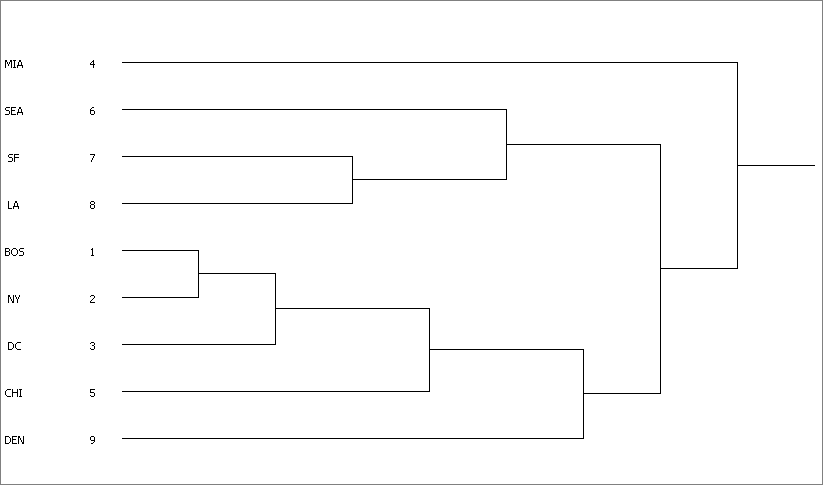
HIERARCHICAL CLUSTERING
M S B C D
I E S L O N D H E
A A F A S Y C I N
Level 4 6 7 8 1 2 3 5 9
----- - - - - - - - - -
206 . . . . XXX . . .
233 . . . . XXXXX . .
379 . . XXX XXXXX . .
671 . . XXX XXXXXXX .
808 . XXXXX XXXXXXX .
996 . XXXXX XXXXXXXXX
1059 . XXXXXXXXXXXXXXX
1075 XXXXXXXXXXXXXXXXX
Measures of cluster adequacy
1 2 3 4 5 6 7
------ ------ ------ ------ ------ ------ ------
1 Eta -0.284 -0.480 -0.554 -0.657 -0.711 -0.687 -0.151
2 Q -0.133 -0.163 -0.188 -0.203 -0.240 -0.214 -0.033
3 Q-prime -0.152 -0.190 -0.226 -0.254 -0.320 -0.322 -0.065
4 E-I 0.994 0.973 0.961 0.884 0.824 0.625 -0.490
Size of each cluster, expressed as a proportion of the total population clustered
1 2 3 4 5 6 7 8
----- ----- ----- ----- ----- ----- ----- -----
1 CL1 0.222 0.333 0.333 0.111 0.111 0.111 0.111 1.000
2 CL2 0.111 0.111 0.111 0.444 0.444 0.333 0.889
3 CL3 0.111 0.111 0.111 0.111 0.333 0.556
4 CL4 0.111 0.111 0.111 0.222 0.111
5 CL5 0.111 0.111 0.222 0.111
6 CL6 0.111 0.111 0.111
7 CL7 0.111 0.111
8 CL8 0.111
Actor-by-Partition indicator matrix saved as dataset Part
----------------------------------------
Running time: 00:00:01
Output generated: 21 11 16 09:10:06
UCINET 6.614 Copyright (c) 1992-2016 Analytic Technologies
E.g. 1
- cities2.csv
0 206 429 1504 963 2976 3095 2979 1949 206 0 233 1308 802 2815 2934 2786 1771 429 233 0 1075 671 2684 2799 2631 1616 1504 1308 1075 0 1329 3273 3053 2687 2037 963 802 671 1329 0 2013 2142 2054 996 2976 2815 2684 3273 2013 0 808 1131 1307 3095 2934 2799 3053 2142 808 0 379 1235 2979 2786 2631 2687 2054 1131 379 0 1059 1949 1771 1616 2037 996 1307 1235 1059 0
# Prepare Data
setwd(“d:/rdata”)
mydata ← read.csv(“cities.csv”)
mydata ← na.omit(mydata) # listwise deletion of missing
mydata ← scale(mydata) # standardize variables