This is an old revision of the document!
Table of Contents
Partial and semi-partial correlation
references
The Elements of Statistical Learning or local copy
Simple explanation of the below procedures is like this:
- Separately regress Y and X1 against X2, that is,
- regress Y against X2 AND
- regress X1 against X2.
- Regress the Y residuals against the X1 residuals.
In the below example,
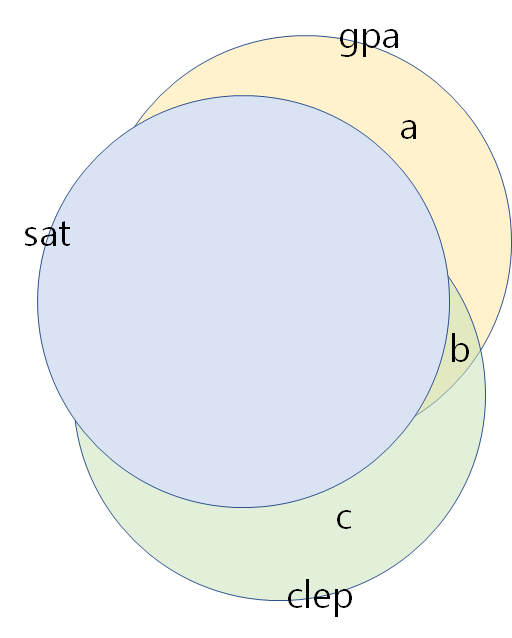
- regress gpa against sat (and get residuals of gpa = a + b)
- regress clep against sat (and get residuals of clep = b + c)
- regress the gpa residuals against clep residuals. (
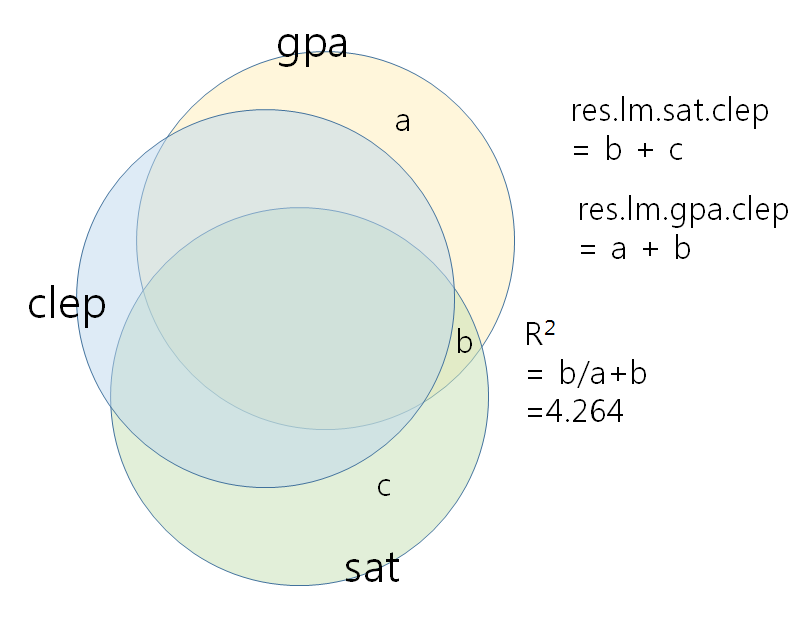
lm(a+b~b+c)) - In this case, $r^{2} = \displaystyle \frac{b}{(a+b)}$ and $b$ is very small.
Take a close look at the right graph, especially, the b areas although clep's is significantly explains gpa (before controlling sat).
For more, see https://stats.stackexchange.com/questions/28474/how-can-adding-a-2nd-iv-make-the-1st-iv-significant
- sat, clep이 각각 (gpa에 대한) regression에 사용되었을 때에 이 둘의 영향력이 나타난다. 그러나, 이 둘을 동시에 같이 사용했을 때에는 sat의 영향력이 사라지게 된다. 따라서 clep에 대해 gpa를 regression 했을 때를 제어하는 것을 보여주기로 한다.
- lm.gpa.clepsat 를 구해서 sat의 영향력이 사라지는 것을 본다.
그리고, res.lm.gpa.clep을 종속변인으로 하고 sat를 독립변인으로 한 영향력을 보면 clep을 제어한 후 sat만의 영향력을 보는 것이 된다 (pcor.test(gpa,sat,clep, data=tests)와 동일해야)- sat에는 clep과 관련이 있는 (상관관계) 부분이 포함되기에 바로 위의 것은 이루어질 수 없다.
- 만약에 독립변인인 sat와 clep이 orthogonal하다면 (즉, 상관관계가 0이라면), 스트라이크 아웃된 부분이 가능하겠지만 그렇지 않기에 sat에서 clep의 부분을 제거한 부분을 구해서 즉, res.lm.sat.clep을 구해서 res.lm.gpa.clep와의 상관관계를 본다.
Partial cor
please refer to the page: http://faculty.cas.usf.edu/mbrannick/regression/Partial.html
tests_cor.csv
| Person | SAT-Q | CLEP | Math GPA |
| 1 | 500 | 30 | 2.8 |
| 2 | 550 | 32 | 3 |
| 3 | 450 | 28 | 2.9 |
| 4 | 400 | 25 | 2.8 |
| 5 | 600 | 32 | 3.3 |
| 6 | 650 | 38 | 3.3 |
| 7 | 700 | 39 | 3.5 |
| 8 | 550 | 38 | 3.7 |
| 9 | 650 | 35 | 3.4 |
| 10 | 550 | 31 | 2.9 |
# import test score data "tests_cor.csv"
tests <- read.csv("http://commres.net/wiki/_media/r/tests_cor.csv")
colnames(tests) <- c("ser", "sat", "clep", "gpa")
tests <- subset(tests, select=c("sat", "clep", "gpa"))
attach(tests)
cors <- cor(tests)
round(cors,3)
sat clep gpa sat 1.000 0.875 0.718 clep 0.875 1.000 0.876 gpa 0.718 0.876 1.000 >
regression gpa against sat
> lm.gpa.sat <- lm(gpa ~ sat, data = tests)
> summary(lm.gpa.sat)
Call:
lm(formula = gpa ~ sat, data = tests)
Residuals:
Min 1Q Median 3Q Max
-0.23544 -0.12184 0.00316 0.02943 0.56456
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.7848101 0.4771715 3.740 0.0057 **
sat 0.0024557 0.0008416 2.918 0.0193 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.2365 on 8 degrees of freedom
Multiple R-squared: 0.5156, Adjusted R-squared: 0.455
F-statistic: 8.515 on 1 and 8 DF, p-value: 0.01935
>
>
linear model
y hat = 0.0024 X + 1.7848
gpa hat = 0.0024 sat + 1.7848
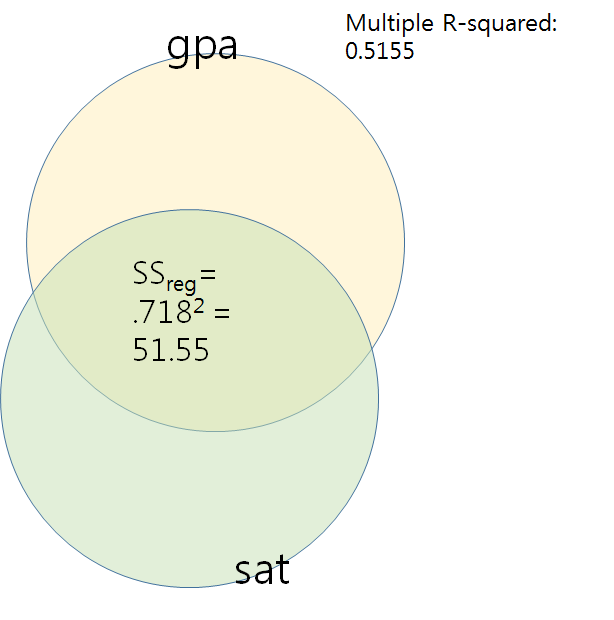
Note that .718 = correlation coefficient of sat and gpa.
> sqrt(0.5156) [1] 0.7180529
Collect
- sat,
- gpa,
- predicted value (y hat),
- residuals (error)
And see correlation among themselves.
> cor.gpa.sat <- as.data.frame(cbind(sat, gpa, lm.gpa.sat$fitted.values, lm.gpa.sat$residuals))
> colnames(cor.gpa.sat) <- c("sat", "gpa", "pred", "resid")
> round(cor.gpa.sat,5)
sat gpa pred resid
1 500 2.8 3.01266 -0.21266
2 550 3.0 3.13544 -0.13544
3 450 2.9 2.88987 0.01013
4 400 2.8 2.76709 0.03291
5 600 3.3 3.25823 0.04177
6 650 3.3 3.38101 -0.08101
7 700 3.5 3.50380 -0.00380
8 550 3.7 3.13544 0.56456
9 650 3.4 3.38101 0.01899
10 550 2.9 3.13544 -0.23544
>
round(cor(cor.gpa.sat),4)
sat gpa pred resid
sat 1.000 0.718 1.000 0.000
gpa 0.718 1.000 0.718 0.696
pred 1.000 0.718 1.000 0.000
resid 0.000 0.696 0.000 1.000
>
Note that
- r (sat and gpa) = .718 (sqrt(r2)=0.5156)
- r (sat and pred) = 1. In other words, predicted values (y hats) are the linear function of x (sat) values (
y hat = 0.0024 X + 1.7848). - r (sat and resid) = 0. residuals are orthogonal to the independent (sat) values.
regression gpa against clep
# import test score data "tests_cor.csv"
tests <- read.csv("http://commres.net/wiki/_media/r/tests_cor.csv")
colnames(tests) <- c("ser", "sat", "clep", "gpa")
tests <- subset(tests, select=c("sat", "clep", "gpa"))
attach(tests)
lm.gpa.clep <- lm(gpa ~ clep, data = tests) summary(lm.gpa.clep)
Call:
lm(formula = gpa ~ clep, data = tests)
Residuals:
Min 1Q Median 3Q Max
-0.190496 -0.141167 -0.002376 0.110847 0.225207
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.17438 0.38946 3.015 0.016676 *
clep 0.06054 0.01177 5.144 0.000881 ***
---
Signif. codes:
0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.1637 on 8 degrees of freedom
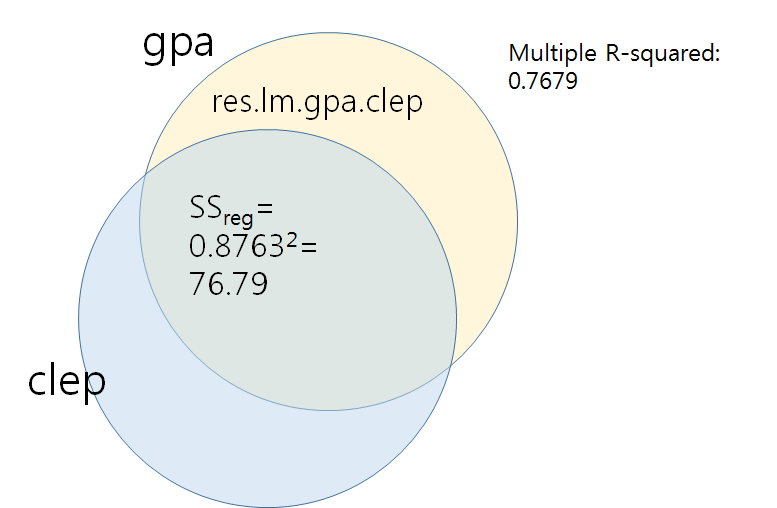
Multiple R-squared: 0.7679, Adjusted R-squared: 0.7388
F-statistic: 26.46 on 1 and 8 DF, p-value: 0.0008808
y hat = 0.06054 * clep + 1.17438
# get residuals res.lm.gpa.clep <- lm.gpa.clep$residuals
# get cor between gpa, sat, pred, and resid from. lm.gpa.clep
cor.gpa.clep <- as.data.frame(cbind(clep, gpa, lm.gpa.clep$fitted.values, lm.gpa.clep$residuals))
colnames(cor.gpa.clep) <- c("clep", "gpa", "pred", "resid")
cor(cor.gpa.clep)
> round(cor(cor.gpa.clep),4)
clep gpa pred resid
clep 1.0000 0.8763 1.0000 0.0000
gpa 0.8763 1.0000 0.8763 0.4818
pred 1.0000 0.8763 1.0000 0.0000
resid 0.0000 0.4818 0.0000 1.0000
>
sat gpa pred resid
sat 1.0000 0.7180 1.0000 0.0000
gpa 0.7180 1.0000 0.7180 0.6960
pred 1.0000 0.7180 1.0000 0.0000
resid 0.0000 0.6960 0.0000 1.0000
>
regression gpa against both celp and sat
lm.gpa.clepsat <- lm(gpa ~ clep + sat, data = tests) summary(lm.gpa.clepsat)
Call:
lm(formula = gpa ~ clep + sat, data = tests)
Residuals:
Min 1Q Median 3Q Max
-0.197888 -0.128974 -0.000528 0.131170 0.226404
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.1607560 0.4081117 2.844 0.0249 *
clep 0.0729294 0.0253799 2.874 0.0239 *
sat -0.0007015 0.0012564 -0.558 0.5940
---
Signif. codes:
0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.1713 on 7 degrees of freedom
Multiple R-squared: 0.7778, Adjusted R-squared: 0.7143
F-statistic: 12.25 on 2 and 7 DF, p-value: 0.005175
>
Multiple R-squared: 0.7778
F (2, 7) = 12.25, p = 0.005157
intercept 1.1607560 p = 0.0249
clep 0.0729294 p = 0.0239
sat 0.0007015 p = 0.5940
One other thing that we could do help determine a pragmatic argument is to regress GPA on both SAT and CLEP at the same time to see what happens. If we do that, we find that R-square for the model is .78, F = 12.25, p < .01. The intercept and b weight for CLEP are both significant, but the b weight for SAT is not significant. The values are
Intercept = 1.16, t=2.844, p < .05CLEP = 0.07, t=2.874, p < .05SATQ = -.0007, t=-0.558, n.s.
In this case, we would conclude that the significant unique predictor is CLEP. Although SAT is highly correlated with GPA, it adds nothing to the prediction equation once the CLEP score is entered. (These data are fictional and the sample size is much too small to run this analysis. It's there for illustration only.)
Now suppose we wanted to argue something a little different. Suppose we had a theory that said that all measures of math achievement share a common explanation, which is math ability. In other words, the reason that various (all) math achievement tests are correlated is that they share the math ability factor. In other words, math ability explains the correlation between achievement tests. In path diagram form, we might represent this something like this:
checking partial cor 1
# get res.lm.clep.sat lm.sat.clep <- lm(sat ~ clep, data = tests) summary(lm.sat.clep)
Call:
lm(formula = sat ~ clep, data = tests)
Residuals:
Min 1Q Median 3Q Max
-101.860 -19.292 1.136 28.306 54.132
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -19.421 114.638 -0.169 0.86967
clep 17.665 3.464 5.100 0.00093 ***
---
Signif. codes:
0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 48.2 on 8 degrees of freedom
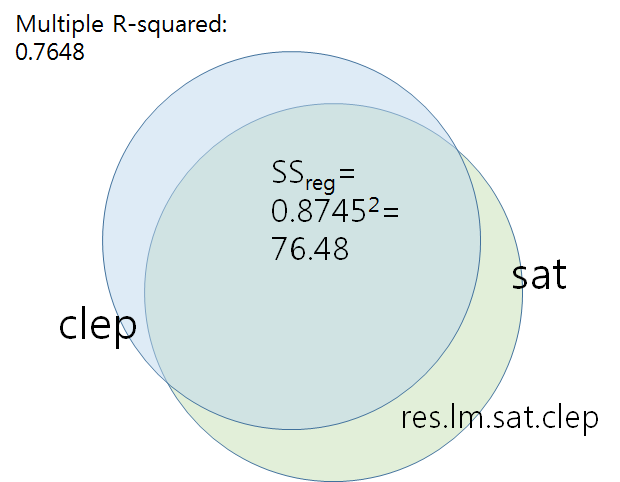
Multiple R-squared: 0.7648, Adjusted R-squared: 0.7353
F-statistic: 26.01 on 1 and 8 DF, p-value: 0.0009303
>
res.lm.sat.clep <- lm.sat.clep$residuals
install.packages("ppcor")
library(ppcor)
pcor.gpa.sat.clep <- lm(res.lm.gpa.clep ~ res.lm.sat.clep) summary(pcor.gpa.sat.clep)
Call:
lm(formula = res.lm.gpa.clep ~ res.lm.sat.clep)
Residuals:
Min 1Q Median 3Q Max
-0.197888 -0.128974 -0.000528 0.131170 0.226404
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.755e-17 5.067e-02 0.000 1.000
res.lm.sat.clep -7.015e-04 1.175e-03 -0.597 0.567
Residual standard error: 0.1602 on 8 degrees of freedom
Multiple R-squared: 0.04264, Adjusted R-squared: -0.07703
F-statistic: 0.3563 on 1 and 8 DF, p-value: 0.5671
> pcor.gpa.sat.clep <- pcor.test(gpa,sat,clep)
> pcor.gpa.sat.clep
estimate p.value statistic n gp Method
1 -0.2064849 0.5940128 -0.55834 10 1 pearson
> pcor.gpa.sat.clep$estimate^2
[1] 0.04263601
>
d <- data.frame(sat=sat, clep=clep, gpa=gpa, res.lm.gpa.clep=res.lm.gpa.clep) plot(d)
Note that the relationship between res.lm.gpa.clep and sat look like negative, which can be confirmed in the lm.gpa.satclep
summary(lm.gpa.satclep)
.
checking partial cor 2
> # import test score data "tests_cor.csv"
> tests <- read.csv("http://commres.net/wiki/_media/r/tests_cor.csv")
> colnames(tests) <- c("ser", "sat", "clep", "gpa")
> tests <- subset(tests, select=c("sat", "clep", "gpa"))
> attach(tests)
> cor(tests)
sat clep gpa
sat 1.0000000 0.8745001 0.7180459
clep 0.8745001 1.0000000 0.8762720
gpa 0.7180459 0.8762720 1.0000000
$$
r_{12.3} = \frac {r_{12} - r_{13} r_{23} } {\sqrt{1-r_{13}^2} \sqrt{1-r_{23}^2}}
$$
(1 = GPA, 2 = CLEP, 3 = SAT)
\begin{eqnarray*} r_{\text{gpaclep.sat}} & = & \frac {r_{\text{gpaclep}} - r_{\text{gpasat}} r_{\text{clepsat}} } {\sqrt{1-r_{\text{gpasat}}^2} \sqrt{1-r_{\text{clepsat}}^2}} \\ & = & \frac {0.8762720 - (0.7180459)(0.8745001)}{\sqrt{1-0.7180459^2} \sqrt{1-0.8745001^2}} \\ & = & .73 \end{eqnarray*}
$$
r_{12.3} = \frac {r_{12} - r_{13} r_{23} } {\sqrt{1-r_{13}^2} \sqrt{1-r_{23}^2}}
$$
(1 = gpa, 2 = sat, 3 = clep)
\begin{eqnarray*} r_{\text{gpasat.clep}} & = & \frac {r_{\text{gpasat}} - r_{\text{gpaclep}} r_{\text{satclep}} } {\sqrt{1-r_{\text{gpaclep}}^2} \sqrt{1-r_{\text{satclep}}^2}} \\ & = & \frac {0.7180459 - (0.8762720)(0.8745001)}{\sqrt{1-0.8762720^2} \sqrt{1-0.8745001^2}} \\ & = & 0.04263585 \end{eqnarray*}
> cor(tests)
sat clep gpa
sat 1.0000000 0.8745001 0.7180459
clep 0.8745001 1.0000000 0.8762720
gpa 0.7180459 0.8762720 1.0000000
> round(cor(tests),4)
sat clep gpa
sat 1.0000 0.8745 0.7180
clep 0.8745 1.0000 0.8763
gpa 0.7180 0.8763 1.0000
> c<- (.7180459-(.8762720*.8745001))
> d <- (sqrt(1-.8762720^2) * sqrt(1-.8745001^2))
> c/d
[1] -0.2064845
> (c/d)^2
[1] 0.04263585
>
Semipartial cor
See also enter page.
> tests <- read.csv("http://commres.net/wiki/_media/r/tests_cor.csv")
> colnames(tests) <- c("ser", "sat", "clep", "gpa")
> tests <- subset(tests, select=c("sat", "clep", "gpa"))
> attach(tests)
> cors <- cor(tests)
> round(cors,3)
sat clep gpa
sat 1.000 0.875 0.718
clep 0.875 1.000 0.876
gpa 0.718 0.876 1.000
> lm.sat.clep <- lm(sat ~ clep, data = tests)
> summary(lm.sat.clep)
Call:
lm(formula = sat ~ clep, data = tests)
Residuals:
Min 1Q Median 3Q Max
-101.860 -19.292 1.136 28.306 54.132
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -19.421 114.638 -0.169 0.86967
clep 17.665 3.464 5.100 0.00093 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 48.2 on 8 degrees of freedom
Multiple R-squared: 0.7648, Adjusted R-squared: 0.7353
F-statistic: 26.01 on 1 and 8 DF, p-value: 0.0009303
> res.lm.sat.clep <- lm.sat.clep$residuals
>
> install.packages("ppcor")
> library(ppcor)
Loading required package: MASS
> # regression test for semipartial correlation (holding clep constant)
> spcor.gpa.sat.clep <- lm(gpa ~ res.lm.sat.clep)
> summary(spcor.gpa.sat.clep)
Call:
lm(formula = gpa ~ res.lm.sat.clep)
Residuals:
Min 1Q Median 3Q Max
-0.3756 -0.2694 -0.0092 0.2514 0.4686
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 3.1600000 0.1069377 29.550 1.86e-09 ***
res.lm.sat.clep -0.0007015 0.0024806 -0.283 0.785
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.3382 on 8 degrees of freedom
Multiple R-squared: 0.009898, Adjusted R-squared: -0.1139
F-statistic: 0.07997 on 1 and 8 DF, p-value: 0.7845
From the above: Multiple R-squared: 0.009898
From the below: spcor.gpa.sat.clep$estimate^2: 0.009897835
> spcor.gpa.sat.clep <- spcor.test(gpa,sat,clep)
> spcor.gpa.sat.clep
estimate p.value statistic n gp Method
1 -0.09948786 0.7989893 -0.2645326 10 1 pearson
> spcor.gpa.sat.clep$estimate^2
[1] 0.009897835
>
e.g.,
In this example, the two IVs are orthogonal to each other (not correlated with each other). Hence, regress res.y.x2 against x1 would not result in any problem.
n <- 32 set.seed(182) u <-matrix(rnorm(2*n), ncol=2) u0 <- cbind(u[,1] - mean(u[,1]), u[,2] - mean(u[,2])) x <- svd(u0)$u eps <- rnorm(n) y <- x %*% c(0.05, 1) + eps * 0.01 x1 <- x[,1] x2 <- x[,2] dset <- data.frame(y,x1,x2) head(dset)
y x1 x2 1 0.2311 -0.42320 0.2536 2 -0.1708 -0.13428 -0.1573 3 0.1617 0.12404 0.1580 4 0.1111 0.10377 0.1214 5 0.2176 0.08796 0.1962 6 0.2054 0.02187 0.1950 >
round(cor(dset), 3)
y x1 x2 y 1.000 0.068 0.996 x1 0.068 1.000 0.000 x2 0.996 0.000 1.000 >
lm.y.x1 <- lm(y ~ x1) summary(lm.y.x1)
Call:
lm(formula = y ~ x1)
Residuals:
Min 1Q Median 3Q Max
-0.3750 -0.1013 -0.0229 0.1402 0.2985
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.00258 0.03242 -0.08 0.94
x1 0.06895 0.18341 0.38 0.71
Residual standard error: 0.183 on 30 degrees of freedom
Multiple R-squared: 0.00469, Adjusted R-squared: -0.0285
F-statistic: 0.141 on 1 and 30 DF, p-value: 0.71
lm.y.x1x2 <- lm(y ~ x1 + x2) summary(lm.y.x1x2)
Call:
lm(formula = y ~ x1 + x2)
Residuals:
Min 1Q Median 3Q Max
-0.026697 -0.004072 0.000732 0.006664 0.017220
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.00258 0.00168 -1.54 0.14
x1 0.06895 0.00949 7.27 5.3e-08 ***
x2 1.00328 0.00949 105.72 < 2e-16 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.00949 on 29 degrees of freedom
Multiple R-squared: 0.997, Adjusted R-squared: 0.997
F-statistic: 5.61e+03 on 2 and 29 DF, p-value: <2e-16
>
….
Std. Error for x1
SSres/n-2 =
Std. Error for x2
….
lm.y.x2 <- lm(y ~ x2) summary(lm.y.x2)
Call:
lm(formula = y ~ x2)
Residuals:
Min 1Q Median 3Q Max
-0.027366 -0.010654 0.002941 0.009922 0.027470
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.002576 0.002770 -0.93 0.36
x2 1.003276 0.015669 64.03 <2e-16 ***
---
Signif. codes:
0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.01567 on 30 degrees of freedom
Multiple R-squared: 0.9927, Adjusted R-squared: 0.9925
F-statistic: 4100 on 1 and 30 DF, p-value: < 2.2e-16
res.lm.y.x2 <- lm.y.x2$resdiduals
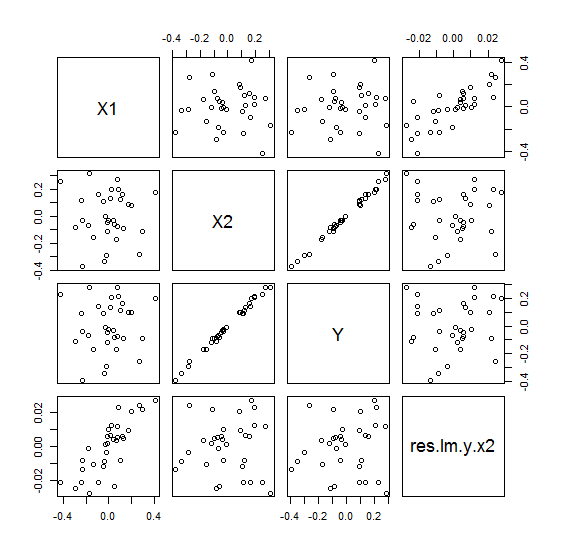
d <- data.frame(X1=x1, X2=x2, Y=y, res.lm.y.x2=res.lm.y.x2) plot(d)
> 1-0.9927 [1] 0.0073
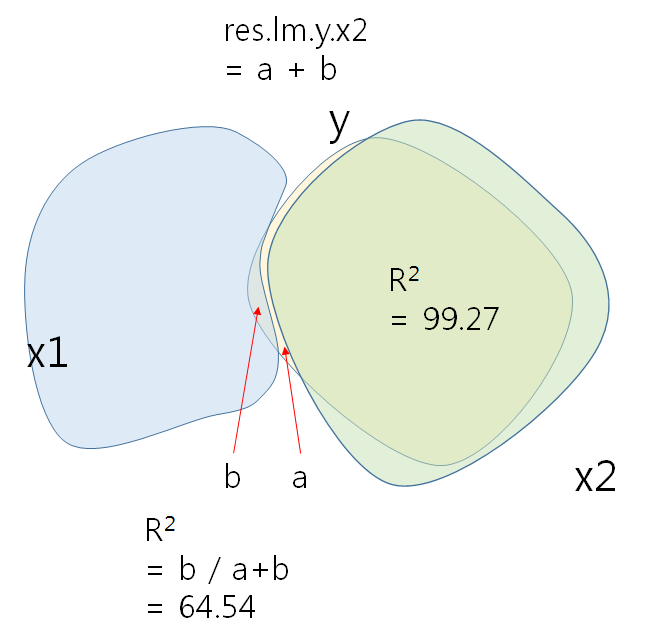
X2이 설명하는 Y분산의 나머지를 (1-R2 = 0.0073) 종속변인으로 하고 x1을 독립변인으로 하여 regression을 하면 figure의 RY축에 해당하는 관계가 나타난다. 특히 RY와 X1과의 관계가 선형적으로 바뀐것은 X1 자체로는 아무런 역할을 하지 못하는 것으로 나타나다가, X2가 개입되고 X2의 영향력으로 설명된 Y부분을 제외한 (제어한, controlling) 나머지에 대한 X1의 설명력이 significant하게 바뀐 결과이다.
> lm.resyx2.x1 <- lm(lm.y.x2$residuals ~ x1)
> summary(lm.resyx2.x1)
Call:
lm(formula = lm.y.x2$residuals ~ x1)
Residuals:
Min 1Q Median 3Q Max
-0.0266967 -0.0040718 0.0007323 0.0066643 0.0172201
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.220e-18 1.649e-03 0.00 1
x1 6.895e-02 9.331e-03 7.39 3.11e-08 ***
---
Signif. codes:
0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.009331 on 30 degrees of freedom
Multiple R-squared: 0.6454, Adjusted R-squared: 0.6336
F-statistic: 54.61 on 1 and 30 DF, p-value: 3.115e-08
>
Actual correlation would look like the below.
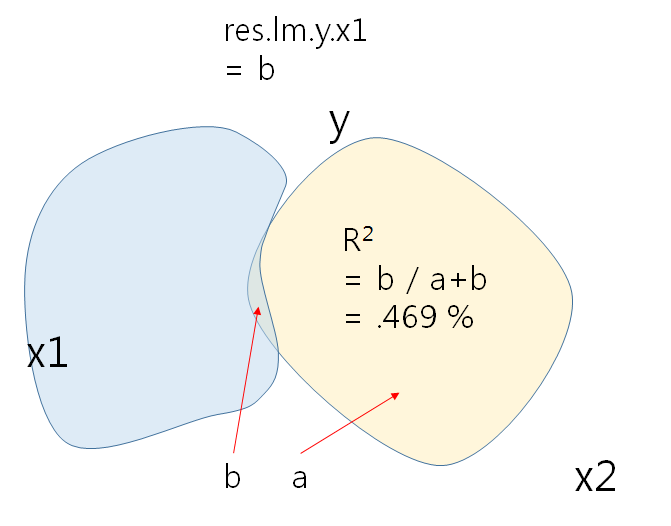
x1의 영향력은 y 총분산에 비해 크지 않다 (b / a + b = .469%)
e.g.
- eg.b.csv
y x1 x2 1.644540 1.063845 .351188 1.785204 1.203146 .200000 -1.36357 -.466514 -.961069 .314549 1.175054 .800000 .317955 .100612 .858597 .970097 2.438904 1.000000 .664388 1.204048 .292670 -.870252 -.993857 -1.89018 1.962192 .587540 -.275352 1.036381 -.110834 -.246448 .007415 -.069234 1.447422 1.634353 .965370 .467095 .219813 .553268 .348095 -.285774 .358621 .166708 1.498758 -2.87971 -1.13757 1.671538 -.310708 .396034 1.462036 .057677 1.401522 -.563266 .904716 -.744522 .297874 .561898 -.929709 -1.54898 -.898084 -.838295
- eg.c.csv
y x1 x2 1.644540 1.063845 .351188 1.785204 -1.20315 .200000 -1.36357 -.466514 -.961069 .314549 1.175054 .800000 .317955 -.100612 .858597 .970097 1.438904 1.000000 .664388 1.204048 .292670 -.870252 -.993857 -1.89018 1.962192 -.587540 -.275352 1.036381 -.110834 -.246448 .007415 -.069234 1.447422 1.634353 .965370 .467095 .219813 .553268 .348095 -.285774 .358621 .166708 1.498758 -2.87971 -1.13757 1.671538 -.810708 .396034 1.462036 -.057677 1.401522 -.563266 .904716 -.744522 .297874 .561898 -.929709 -1.54898 -1.26108 -.838295
e.g. 3,
set.seed(888) # for reproducibility S = rnorm(60, mean=0, sd=1.0) # the Suppressor is normally distributed U = 1.1*S + rnorm(60, mean=0, sd=0.1) # U (unrelated) is Suppressor plus error R <- rnorm(60, mean=0, sd=1.0) # related part; normally distributed OV = U + R # the Other Variable is U plus R Y = R + rnorm(60, mean=0, sd=2) # Y is R plus error
e.g. 4 Using ppcor.test
## install.packages("psych")
## install.packages("ppcor")
library(pysch)
library(ppcor)
## ppcor(v.dv, v.iv, v.ctrl)
options(digits = 4)
SATV <- c(500, 550, 450, 400, 600, 650, 700, 550, 650, 550)
HSGPA <- c(3.0, 3.2, 2.8, 2.5, 3.2, 3.8, 3.9, 3.8, 3.5, 3.1)
FGPA <- c(2.8, 3.0, 2.8, 2.2, 3.3, 3.3, 3.5, 3.7, 3.4, 2.9)
scholar <- data.frame(SATV, HSGPA, FGPA) # collect into a data frame
describe(scholar) # provides descrptive information about each variable
corrs <- cor(scholar) # find the correlations and set them into an object called 'corrs'
corrs # print corrs
pairs(scholar) # pairwise scatterplots
pcor.test(HSGPA,FGPA,SATV)
reg1 <- lm(HSGPA ~ SATV) # run linear regression
resid1 <- resid(reg1) # find the residuals - HSGPA free of SATV
reg2 <- lm(FGPA ~ SATV) # second regression
resid2 <- resid(reg2) # second set of residuals - FGPA free of SATV
cor(resid1, resid2) # correlation of residuals - partial correlation
options(digits = 4) HSGPA <- c(3.0, 3.2, 2.8, 2.5, 3.2, 3.8, 3.9, 3.8, 3.5, 3.1) SATV <- c(500, 550, 450, 400, 600, 650, 700, 550, 650, 550) FGPA <- c(2.8, 3.0, 2.8, 2.2, 3.3, 3.3, 3.5, 3.7, 3.4, 2.9) GREV <- c(600, 670, 540, 800, 750, 820, 830, 670, 690, 600) ##GREV <- c(510, 670, 440, 800, 750, 420, 830, 470, 690, 600) scholar <- data.frame(HSGPA, SATV, FGPA, GREV) # collect into a data frame describe(scholar) # provides descrptive information about each variable corrs <- cor(scholar) # find the correlations and set them into an object called 'corrs' corrs # print corrs pairs(scholar) # pairwise scatterplots pcor.test(HSGPA,FGPA,SATV) reg1 <- lm(HSGPA ~ SATV) # run linear regression resid1 <- resid(reg1) # find the residuals - HSGPA free of SATV reg2 <- lm(FGPA ~ SATV) # second regression resid2 <- resid(reg2) # second set of residuals - FGPA free of SATV cor(resid1, resid2) # correlation of residuals - partial correlation reg12 <- lm(HSGPA ~ GREV) resid12 <- resid(reg12)